Abstract
In the last few years, a renewed interest has been devoted to the origin of life. Nucleic acids have a central role in biogenesis and prebiotic synthesis of ribose, purines and pyrimidines has been described. Particularly attractive is the idea that life on Earth was first founded on simple and autocatalytic RNA molecules (RNA world). RNA duplexes are formed by helical and antiparallel chains, but the rationale for these features is not yet known. An inverted (antiparallel) stacking of purine rings from guanosine and other nucleosides has been found in crystallographic studies. Molecular modeling also supports an inverted conformation, and this preferential stacking occurs when they are aggregated within a clay (montmorillonite) matrix, which is confirmed by spectral studies. Thus, crystallography, spectroscopy and molecular modeling allow proposing a "zipper" model in which antiparallel nucleosides stack within the clay matrix, and then high-energy polyphosphates form auto-intercalated single RNA chains. Unstacking and strand separation result in the shortening and inclination of the ribose-phosphate chains, leading to helicity, antiparallelism and base pairing by H-bonding, with formation of typical RNA structures such as hairpins, cloverleaves, hammerheads, dumbbells, circular viroids and virusoids. A "tetris" model for the prebiotic self-replication mechanism of these simple RNA duplexes, involving the formation of an RNA tetraplex, is also proposed.
Keywords
1. Introduction
Several theories of how life appeared on Earth have inspired controversial and often inaccurate statements, including claims about its extraterrestrial origin. In order to approach this problem, both geometrical and mechanistic rationale are applied in this work to attempt understanding how RNA molecules could have been originated and replicated in a prebiotic environment. To this respect, some old and recent reviews on the origin of life could be consulted[1-10]. Various building blocks of macromolecules were found in meteorites and they could have been delivered to Earth in its infancy from outer space. However, it is currently accepted that all these spatial organic molecules were also formed on Earth as building blocks along the prolonged prebiotic period in our planet.
The period for biogenesis, from the simple organic chemistry scenario to a primitive but already complex cell, seems to be "only" about 0.4 x 109 years[6]. The creation of many different organic molecules under conditions mimicking the prebiotic Earth is now well-established[11-13]. These organic molecules are a wide range of C-, H-, N-, O-containing compounds, aromatic heterocycles, sugars and phosphate esters, which are well known components of coenzymes, biopolymers and particularly, nucleic acids (NAs), namely, RNA.
The attractive "RNA world" concept[5,6,14,15] is now considered the most plausible scenario for the origin of cellular life[16,17]. However, in this hypothetical stage of early life, it remains unknown how a pristine RNA would initially have been synthesized without the aid of protein enzyme catalysis and a previously formed template for replication[18].
An interesting approach about the origin of life is the recently formulated assembly theory[19]. According to this proposal, it is possible to objectively measure the complexity of biomolecules by calculating the minimum number of steps needed to make them from their chemical components, which is quantified by the assembly index[19,20]. However, in the assembly theory, geometrical parameters and molecular shape and size of precursors are not taken into account.
Numerous chemical processes that form organic molecules are known as "prebiotic chemistry", which can use energy from electrical discharges, heating, radioactivity, cosmic, X, γ, UV, visible, and IR radiation, etc., a fact that has been reviewed[10,21,22]. Although no central to the present study, some indications about the origin of organic molecules such as NAs precursors must be given. It is accepted that life arose in a prebiotic soup of chemicals. Organic precursor compounds (heterocyclic bases, sugars, amino acids) were the main components of the primordial soup[2,23], and were also subject of a natural early/primal evolution.
Nitrogenous bases (nucleobases) such as purines (adenine, guanine) and pyrimidines (uracil, thymine, cytosine) arose from polymerization of hydrogen cyanide[6,12]. Pyrimidine nucleosides can be built through a reaction between cyanamide and glycolaldehyde[24]. Nucleobases have been also found in carbonaceous meteorites[25]. Carbohydrates (sugars) originated from autocatalytic polymerization (aldol condensation and enolization) of formaldehyde according the "formose" cascade reactions discovered in 1861 by Butlerow[13]. Specifically, D-ribose and other pentoses result from the addition of glycol-aldehyde to β-hydroxyketone.
The origin of nucleosides involves the formation of a β-N-glycosidic bond between the C1' atom of D-ribose (β-D-ribofuranose) and N9 of purine or N3 of pyrimidine rings[6]. However, the structural basis of the formation of polynucleotides from known precursors remains, at the present, not yet fully understood and therefore, more studies on the organization and assembly of building blocks are still lacking.
The structure, physicochemical properties and biological functions of NAs have been exhaustively studied and reviewed[26-32]. The prebiotic formation of RNA macromolecules is a pristine and highly relevant event in the origin of life, but their non-enzymatic synthesis is scarcely known, and more chemical, mechanistic, and computational studies should be considered to unveil the rationale for the puzzling helical and antiparallel orientation of these biopolymers. To this respect, some conceptual engineering approaches relate with macromolecular properties and processes of assembly and self-assembly of components. They have been introduced in theoretical and applied studies, examples being new research fields such as chemical, molecular, genetic, genomic, tissue, material, and metabolic engineering[33,34].
Regarding strandedness of NAs, there are two main types according to the number of polynucleotide chains: single-stranded (ss), and double-stranded (ds) forms. Although ss-NAs are organized as a simple 3'-5' orientated polymer, it is rather surprising that in ds-NAs the orientation of ribose-phosphate chains is just opposed, namely, antiparallel. Figure 1 illustrates this fact, as well as the general organization of a ds-polyribonucleotide chain.
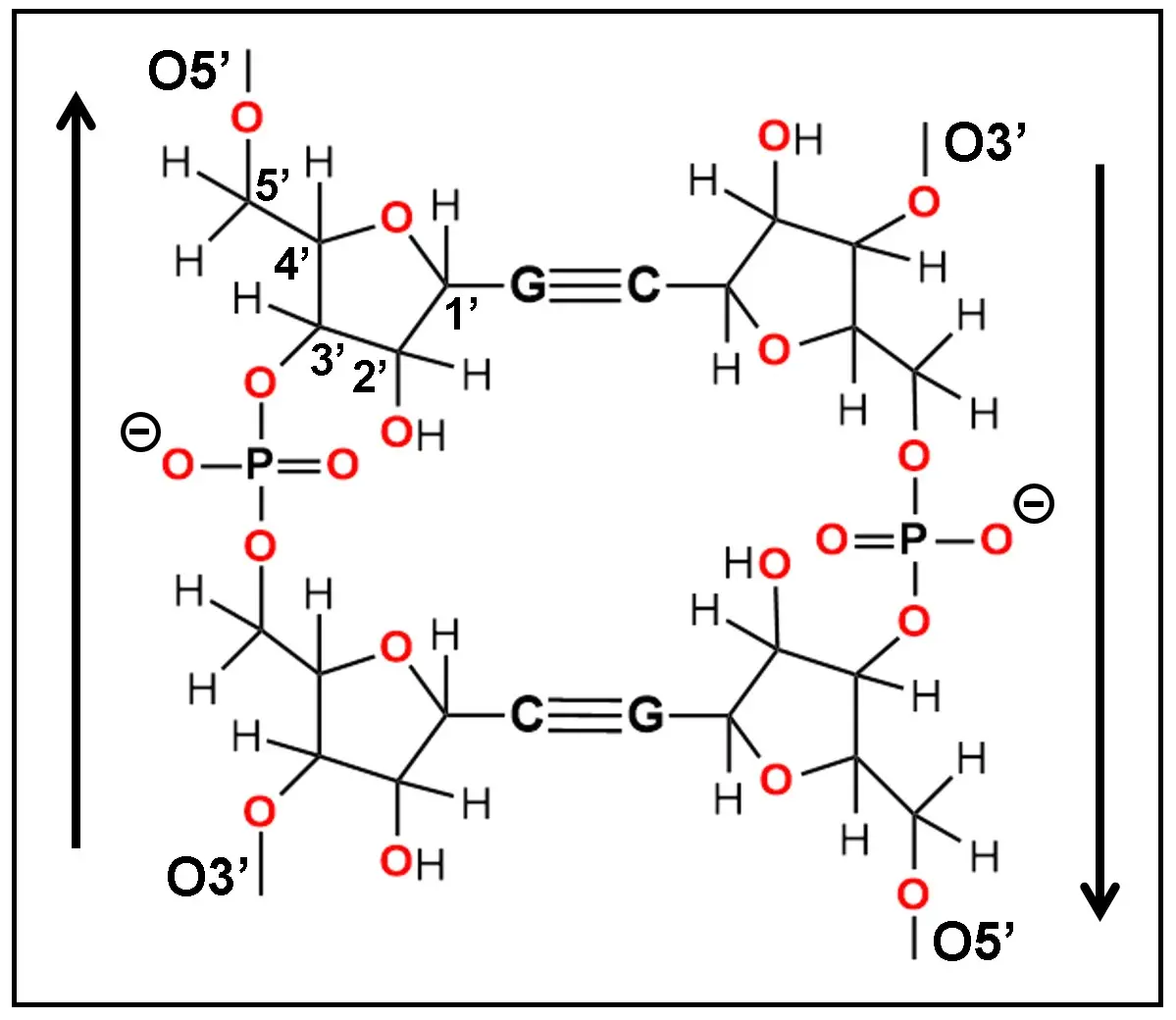
Figure 1. Primary chemical structure of the ribose-phosphate chains from a ds-RNA segment showing their antiparallel (3'-5' and 5'-3') orientation indicated by vertical arrows. Oxygen atoms are in red and the numbering of carbon atoms from ribose is shown. Observe the inverted position of the pentagonal furanose ring from both chains. G and C are guanine and cytosine bases paired by three WC H-bonds. WC: Watson-Crick.
Oligoribonucleotides containing guanine (G) and cytosine (C) nitrogenous bases are mainly used here as a case study to introduce innovative conceptions about the self-assembly of nucleic acids precursors and to substantiate the present model. For illustrative purposes and easier understanding of the building process, the general organization, right-handed helicity, 3' → 5' orientation and antiparallelism of typical ss- and ds-RNA segments can be clearly observed in Figure 2A,B.
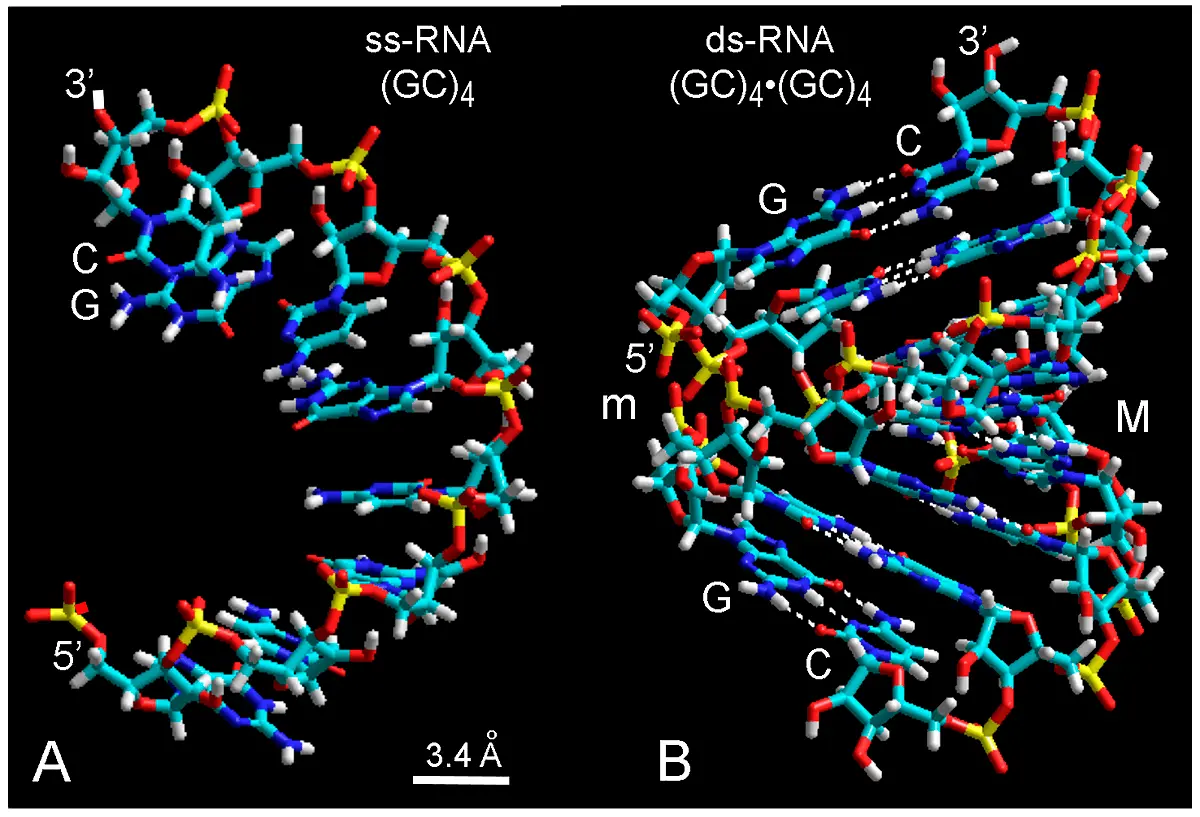
Figure 2. Molecular structure (lateral view) of ss-RNA (A) and ds-RNA (B) modeled with 3'-endo ribose conformation, using the databases of G and C nucleotides and tube rendering. In (B), white dashed lines are H-bonds between base pairs with high tilt angle (about 10º-20º[29]). The shallow minor groove (m) and deep major groove (M) are shown. In both cases, the helical pattern of chains with stacked and paired bases results apparent. The 3' and 5' ends of ribose are indicated. Color code of elements is C: cyan, O: red, N: blue, H: white, P: yellow.
But how have these NAs become so complex? When we observe the intriguing structure of ss- and ds-RNAs, it is certainly difficult to imagine the origin of these biopolymers in the absence of template, enzymatic catalysis and metabolism. Under these conditions, it seems logical to assume that specific chemical geometry and reactivity are the unique factors involved in self-assembly processes that occurred to form complex NA polymers. In this work, using a combination of crystallographic data, chemical formulations, spectroscopy, and molecular computational studies, a mechanistic model for explaining the prebiotic origin of the RNA structure is proposed. Early ideas were inspired on the antiparallel (inverted) stacking of purine and pyrimidine nucleosides[35,36], as well as stacking of aromatic compounds in a clay (montmorillonite) matrix[37,38]. To the extent of the author's knowledge, the proposal that both helicity and antiparallelism originated in the inverted stacking of bases followed by formation of a polyribonucleotide chain within a clay matrix, has been not yet reported.
Taking into account the easy stacking of aromatic heterocyclic compounds within clays, it can be expected that any nucleoside can form inclusion complexes within a suitable montmorillonite cage. This is the main rationale for the present model. First, nucleoside dimers stack within the aqueous interlayer of the clay, and then phosphate bridges and polyribonucleotide chains are formed. This model is therefore based on a clay-induced, self-assembled stacking of nucleosides followed by phosphate linking, and accounts for the origin of helicity and antiparallelism of NAs duplexes. Facts and consequences of this process will be further described in a coordinated way.
2. Methodological Aspects
2.1 Molecular modeling
To substantiate molecular structures and events, simple molecular modeling methods were performed following previous studies to generate computer-drawn skeletal (wire), ball-and-stick, and space-filling (atomic volume) models[39-41]. More accurate methods such as density functional theory to calculate mechanical properties of peptide and protein are complex and expensive[42], and they were not applied. Chemical structures with formal double bonds were drawn with ChemDraw Ultra vl2.0 software. The Dtmm87 molecular modeling program was used in some cases for geometry optimization[43,44]. Geometry optimization and energy minimization were carried out with the friendly HyperChem v7 and v8.0.10 software, using databases for RNA building with the corresponding conformation for preformed nucleotides. Molecular mechanics force field (MM+) was applied for geometry optimization, whereas semi-empirical PM3 Polak-Riviere conjugate gradient and extended-Hückel method for energy minimization and calculation of molecular orbitals (MOs) were used in modeling studies. Geometry optimization and energy minimization of modeled RNA structures were recorded when energy (E) converged at either 0.1, 1, or 10 kcal/(Å mol).
Chemical parameters were chosen according the most classical and accepted values that characterize RNA duplexes. Specific conformations for nucleotides in A-type RNA with 11 base pairs per helix turn were pentose pucker: C3' -endo ; N-C1' glycosidic angle: anti, and torsional angle C4'-C5': gauche-gauche (+ SC)[29,45]. Stereochemical models of RNA duplex r(AU)30 · r(AU)30 and the corresponding RNA tetraplex were automatically modeled from the database set of Hyper Chem v7 menu, using typical parameters for a right-handed A-RNA helix, with C3'- endo and rA and rU bases in anti -conformation.
The ground and excited states of auto-intercalated and antiparallel bis(GC)4 oligomer correspond to different π-electron MOs. The Highest-Occupied (HOMO), and Lowest Unoccupied (LUMO) Molecular Orbital states represent the energy levels of the ground and excited state of the molecule, respectively[46]. In HOMO - d and LUMO + d, "d" is a value from 0 to the maximum energy level, and the HOMO-LUMO separation corresponds to the forbidden Fermi's energy gap (Eg). When electronic energy levels are highly compacted, HOMO and LUMO states form continuum bands, which correspond to the well-known semiconductor-like valence band and conduction band, respectively. Colors of π-electron orbital lobes denote orbital phases. Fused lobes with the same color are in-phase, and those with isolated (unfused) lobes are outof-phase. The analysis of MOs is especially suitable to reveal the coupling of LUMO states in stacked molecules, which expands the excited π-electron resonance along the vertical stacking axis[39-41].
2.2 Absorption and fluorescence spectroscopy
To confirm the stacking of nucleosides within a clay matrix, spectroscopic studies were performed using 9 x 10-5 M guanosine (Sigma-Aldrich, St. Louis, MO, USA) in distilled water at pH 6 and room temperature (22 °C). Guanosine solutions were employed either alone or in the presence of a colloidal suspension (0.2 mg/mL) of montmorillonite (Hotaka Brand, Hojun Hining Co., Tokyo, Japan) as a cosolute matrix. Absorption and emission measurements were made with a Perkin-Elmer UV-vis 551-S spectrophotometer and a Perkin-Elmer 650-10S fluorescence spectrophotometer, respectively (Norwalk, CT, USA). The later instrument was equipped with a 150-W xenon lamp, two grating monochromators, the R372F photomultiplier detector and a 056 recorder.
Fluorescence analysis was made under 260 nm-excitation, with sensitivity range 0.1, mode switch and gain selector in normal position, and a 10-nm bandpass for the excitation and emission slits. The absorption and emission spectra of experimental and control solutions containing the clay alone (subtracted from experimental spectra) were recorded from the supernatant solutions after sedimentation of samples for 48 h. The Raman scattering of the cosolute and aqueous solvent was subtracted from emission spectra.
3. Nucleoside Stacking
H-bonded base pairs can display two kinds of twofold symmetry, depending on the orientation of the glycosyl C1'-N bonds, which give parallel or antiparallel chains[29]. Aggregation of aromatic compound by stacking of molecules is a well-known process. In the case of the interaction of heterocyclic NA bases, early studies showed that in crystals of both guanosine and inosine nucleosides the bases adopt a typical stacking, namely, an inverted configuration[35,47] (Figure 3). In contrast, stacking of the base alone (e.g., guanine) generally present a non-inverted (normal) configuration.
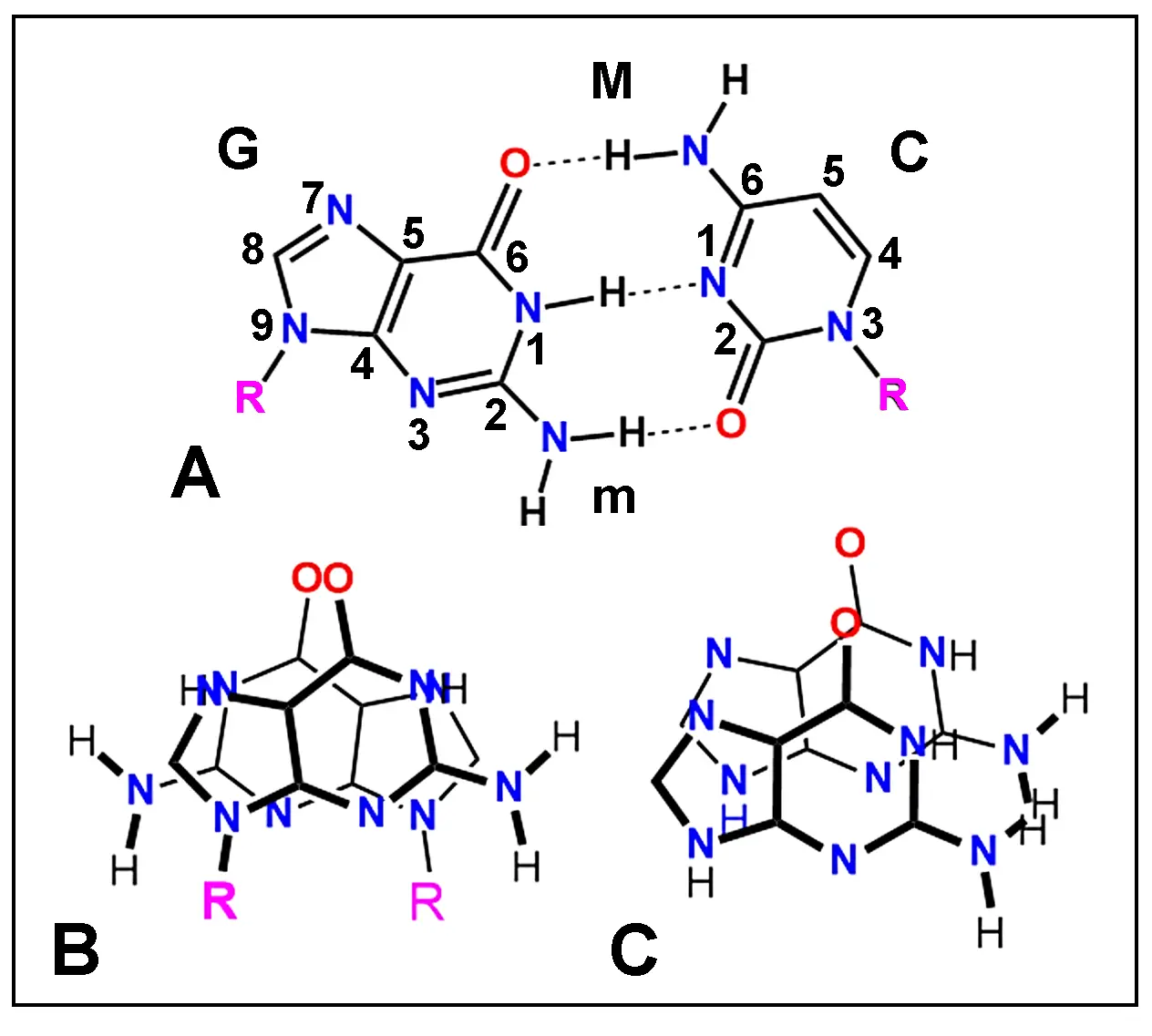
Figure 3. (A) Chemical structure of paired guanosine (G, left) and cytidine (C, right) nucleosides showing atom numbering and H-bonding (dashed lines). In ds-NAs, this pairing pattern results in the formation of two different helical channels, major (M) and minor (m) grooves. (B, C) Crystallographic structure of stacked guanosine (B) and guanine (C) molecules. Observe the inverted (antiparallel) stacking of guanosine (B) and the normal (parallel) stacking of guanine (C). R: ribose.
Molecular mechanics approaches have been widely used in studies involving interaction and docking between chemical compounds[44,48]. In the present case, both guanosine and cytidine do stack closely as observed after geometry optimization of these nucleosides (Figure 4).
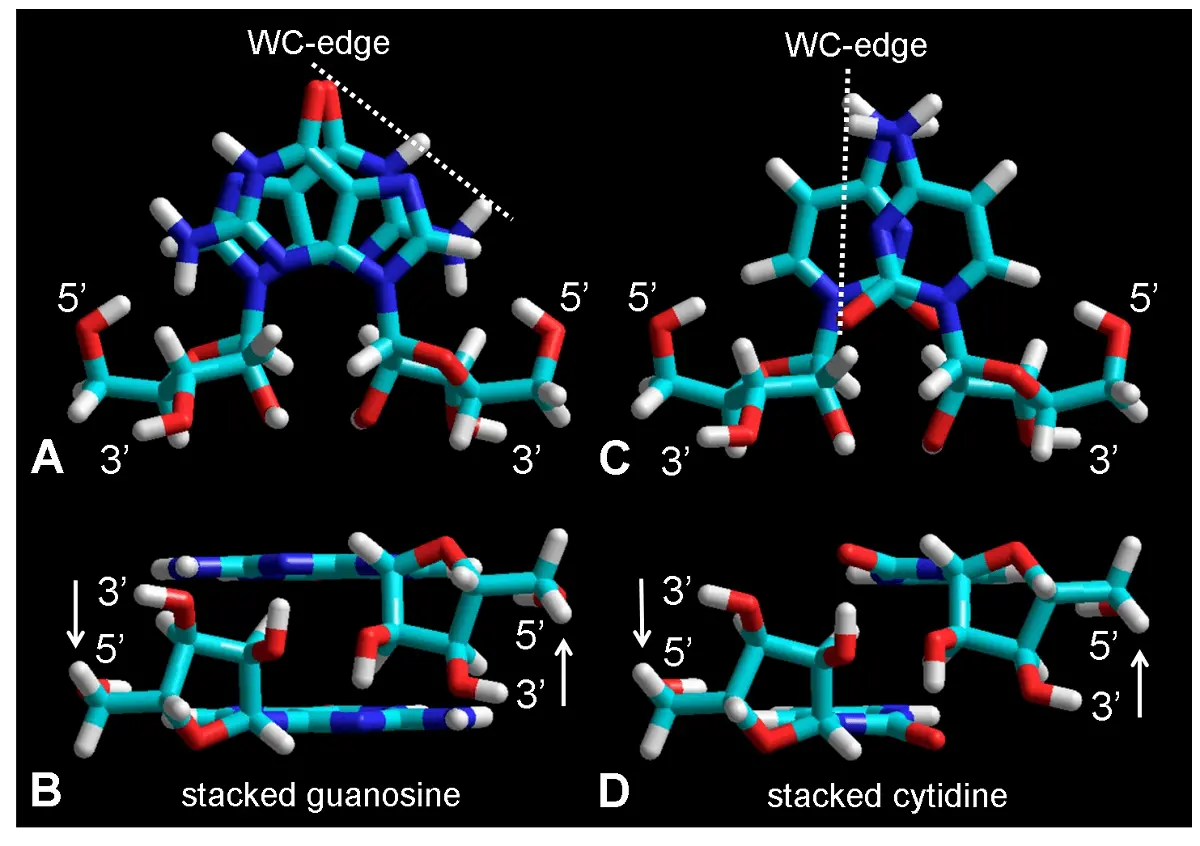
Figure 4. Molecular structure of stacked guanosine (A, B) and cytidine (C, D) dimers shown as tube rendering from frontal (A, C) and lateral (B, D) views. The 3' and 5' atoms from riboses are indicated, as well as the Watson-Crick edges (dashed lines) for possible base pairing by H-bonding. Observe the large and small overlapping between guanine and cytosine bases, respectively, and the antiparallel orientation of the 3' and 5' positions in both nucleosides. Color code of elements is C: cyan, O: red, N: blue, H: white, P: yellow. WC: Watson-Crick.
Stacking of other purine (inosine, adenine) and pyrimidine (uracil, cytosine, thymine) nucleosides also occur[29,49,50], in most cases adopting a recurring inverted orientation. In single crystals, two types of A-A, G-G, T-T and C-C stacking patterns, normal and inverted, are energetically almost equally favorable[36], and normal stacking are often found[47,51]. A substantial contribution into the stability of the secondary structure of polynucleotides is just provided by stacking of bases[26,29,30,50]. Anyway, although normal stacking patterns could also form guanosine dimers in the clay cage, the best proof-of-concept for the present model is that the inverted base stacking is the only configuration that can explain the antiparallelism of ds-NAs.
To substantiate the inverted (antiparallel) stacking of nucleosides in comparison with other possible parallel geometries, calculations of the free energy for different types of stacked dimers were performed. Figure 5 shows that the preferential minimum of free energy for guanosine stacking (E = 51.07 kcal/mol) is precisely that which occurs in the crystallographic study of the inverted guanosine dimer[35].
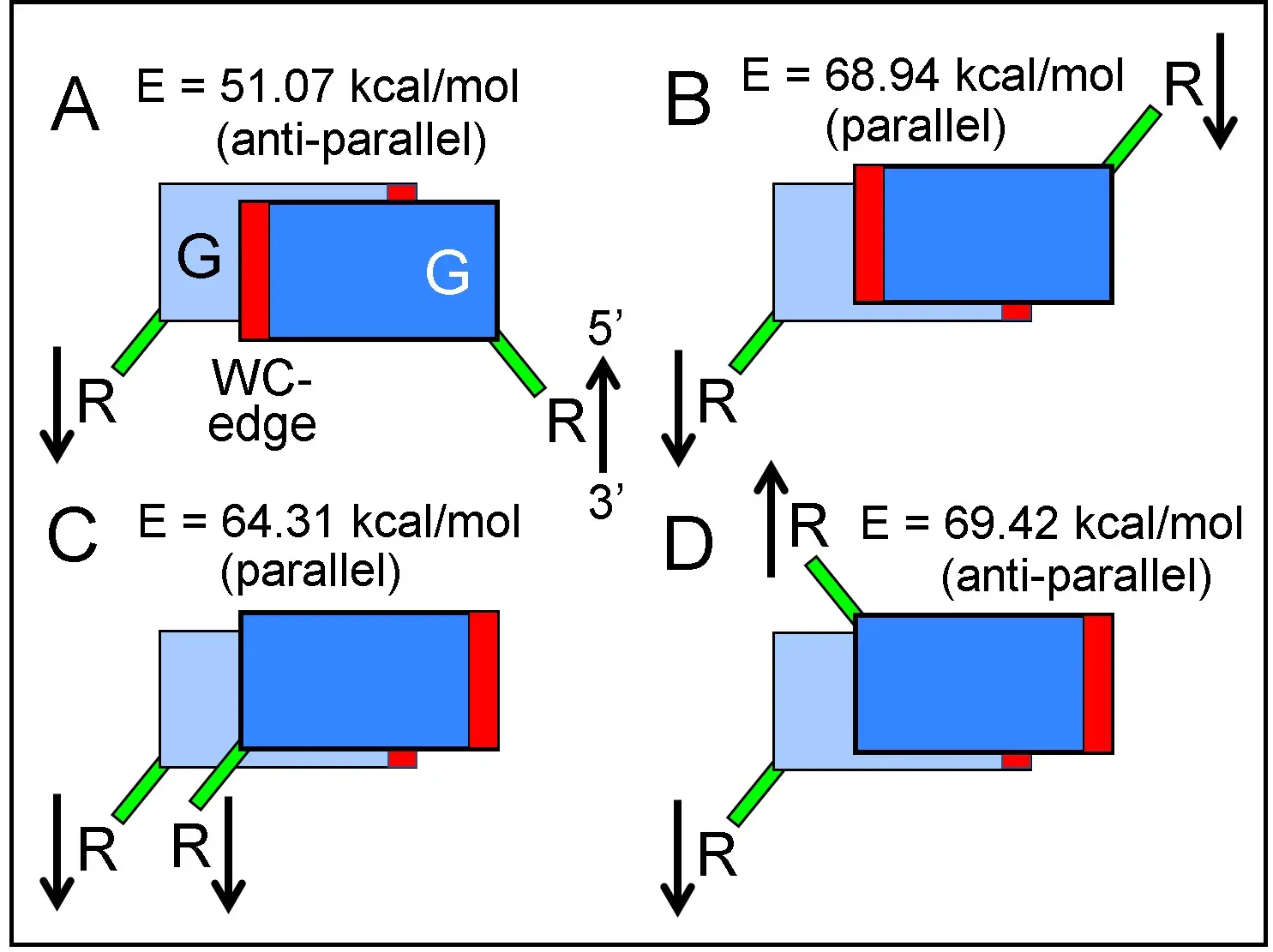
Figure 5. Free energy values calculated for guanosine stacked dimers with different orientations. (A) and (D): Inverted stacking, antiparallel arrows. (B) and (C): normal (non-inverted) stacking, parallel arrows. R: ribose, G: guanine. Green bars: N-glycosidic bonds between bases and riboses. Red bars indicate the position of WC H-bonding edges, in this scheme without true angular position. Observe that a rotation of 180º around perpendicular axis transforms (B) and (C) into an (A) conformation. WC: Watson-Crick.
4. Formation of Nucleotides
Due to auto-intercalation, there is a separation of 6.8 Å between unpaired bases from the same chain, instead of the 3.4 Å distance between paired bases in ds-NAs. It is noteworthy that the inverted base stacking of nucleosides is the most relevant geometric feature of this model, in which purine-purine, pyrimidine-pyrimidine, and purine-pyrimidine stacking can occur. Phosphate linking with high-energy pyrophosphate, tri- or polyphosphates occurs between ribose hydroxyls and PO4 due to the easy nucleophilic attack of O3' and O5' (2.70 Å apart) toward the P atom. This process can take place within the clay matrix due the appropriate distances between the adjacent hydroxyls of riboses, giving nucleotides and then auto-intercalated polynucleotide chains, which are the most characteristic intermediate for RNA formation (Figure 6).
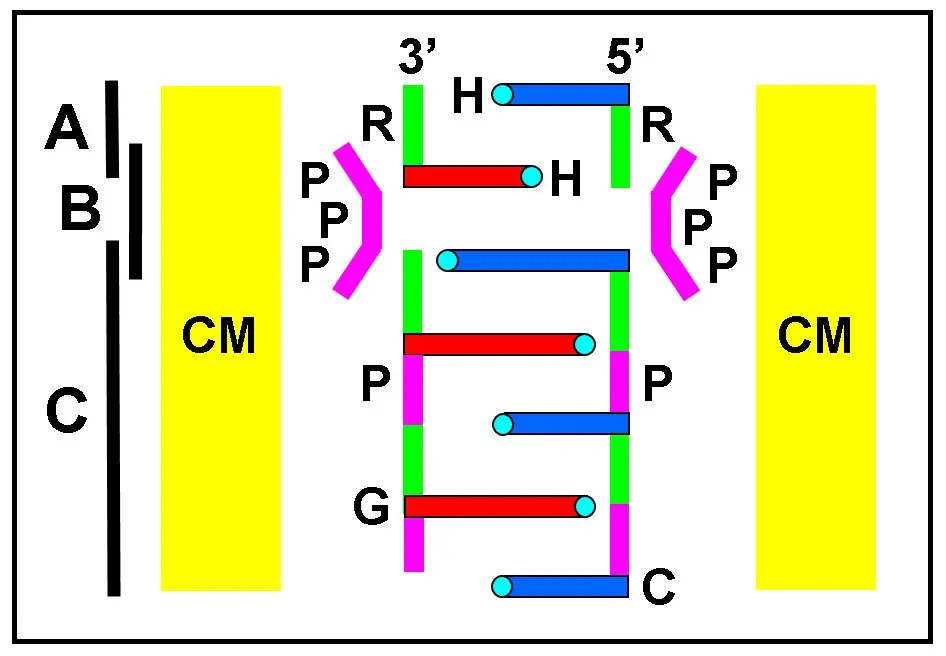
Figure 6. Schematic drawing of the transition steps (vertical black lines) involving (A) stacked nucleoside dimers; (B) phosphate bridging through high-energy triphosphates (PPP), forming (C) auto-intercalated polyribonucleotide chains. All these reactions could develop within the CM of montmorillonite. Horizontal bars are guanine (G, long) and cytosine (C, short) bases. H-bonding groups are small cyan circles. Ribose (R, green) and phosphate (P, violet) are indicated. Bases with identical colors belong to the same chain. CM: clay matrix.
Phosphate-diester bridging with formation of nucleotides and polynucleotides takes place by the nucleophilic attack of O3' and O5' from ribose to the central P from polyphosphate anions. The formation of a phosphate-ester bond utilizes the unique phenomenon of the high free energy of pentavalent phosphorous by breaking the P ~ O bond[21,52], the symbol ~ denoting high-energy bonds. Phosphate esters between riboses are thus formed by the energy-rich bonds of polyphosphate anhydrides (-P ~ O ~ P ~ O ~ P-)[53], giving H2O from the phosphate -OH and from hydrogens of the O3' and O5' hydroxyl. Interestingly, inorganic polyphosphates (polyP) have a great amount of high-energy phosphate bonds (ΔG; Gibbs free energy)[54,55], which can be used in the formation of phosphodiester O3'-P-O5' linkage between riboses. A likely mechanism for the possible formation of phosphate bridges between riboses is graphically shown in Figure 7.
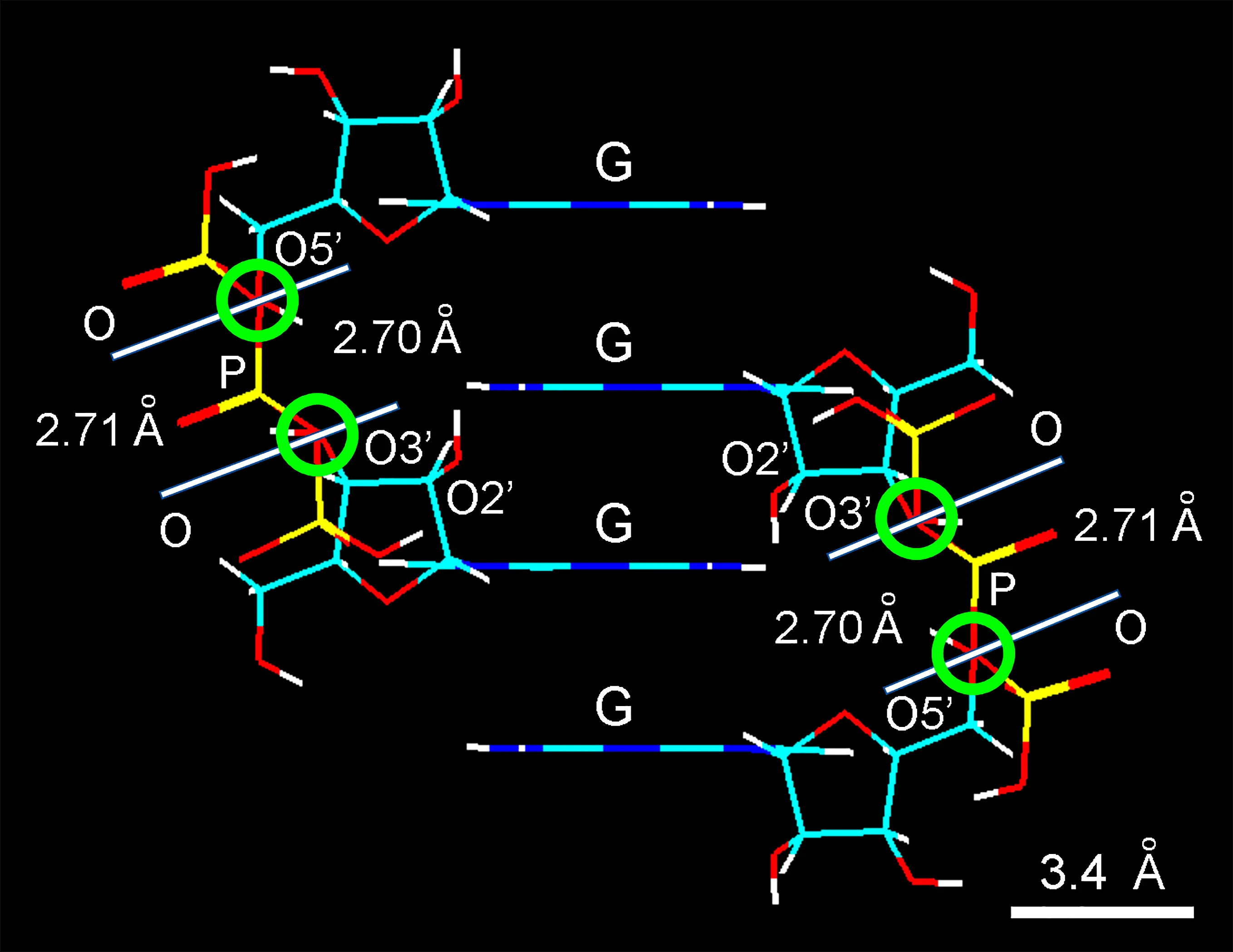
Figure 7. Wire molecular model of two auto-intercalated guanosine dimers showing the close correspondence in the length from O5' and O3' atoms of neighboring nucleosides (2.70 Å) and the linear distance between two O atoms of the central phosphate group (2.71 Å) from the properly inclined triphosphate anion. In each case, a white line indicates the separation between both red-overlapped O atoms (green encircled lines). G: stacked guanine bases. Observe the longer distance between the phosphate O and the ribose O2' to form a suitable diester link. Color code of elements is C: cyan, O: red, N: blue, H: white, P: yellow.
Molecular modeling shows that phosphate is the best group for linking neighboring nucleosides. This reaction mechanism easily accounts for the formation of phosphate bridges and nucleotide polymerization. The energy release by high-energy pyrophosphate hydrolysis (ΔG = -31.8 kJ/mol) is quite sufficient to phosphorylate a ribose hydroxyl (ΔG = -12.6 kJ/mol). In addition, a catalytic activity of specific metal cations (e.g., Mg2+, Mn2+, Zn2+) on clay scaffolding (surfaces or sheet edges) can also promote the formation of phosphate diesters between riboses[56,57]. It is worth to note that DNA and RNA polymerases also contain tightly bound Zn2+ in the active sites[58].
Binding negatively charged compounds occurs easily at the broken edges of montmorillonite sheets. For example, pyrophosphate ions bind selectively at these edges[2]. Interestingly, Ferris[59] mentioned the role of montmorillonite on the assembly of nucleotides to form short RNA polymers. Thus, montmorillonite appears as a prebiotic cage catalyst for the formation of the phosphodiester bonds of RNA[60,61].
In addition to polyP, there are other good phosphorylating agents with high-energy phosphates that could be used in the prebiotic scenario. Examples are acetyl-phosphate, imidoyl-phosphate (from the dihydro-nicotinamide ring of NADH2), diamido-phosphate (DAP) and carbamoyl-phosphate[62-64]. The spontaneous synthesis of carbamoyl-phosphate from cyanate or isocyanic acid and inorganic phosphate allows an easy formation of a carrier of energetic phosphate in the prebiotic world[65]. Imidazoles are also readily formed from simple precursors under prebiotic conditions. Imidazole derivatives (obtained from sugar and ammonia) react with high-energy phosphate to give N-phosphoryl-imidazoles[66], which can be used for the formation of phosphate-diester bridges between riboses in ss-RNA. Likewise, N-cyano-imidazole, or carbodiimides can activate phosphates for attack by a hydroxyl group[67].
Likewise, nucleoside-5'-phosphoryl-imidazoles are suitable for non-enzymatic, untemplated RNA synthesis. The use of activated nucleotide precursors containing phosphoryl-2-methyl- or 2-amino-imidazole groups is an improved way to form polynucleotides[66,68,69]. Taking into account the possibility of easy synthesis of phosphoryl-imidazole derivatives under prebiotic conditions, it is logical to assume that, like pyrophosphate, the simple reactive compound phosphoryl-N,N'-diimidazole[70] could also form a phosphate link between riboses based on the nucleophilic attack of O3' and O5' to the P atom. In spite of hydrolysis, phosphate diesters are rather unreactive compounds[71], which ensures the stability of polynucleotide chains.
5. The Clay Matrix
Since long ago, it is known that the mineral montmorillonite (a very common geological material), is a three-layered clay composed of external sheets of tetrahedral (T) silicates and internal octahedral (O) alumina groups[72,73]. The crystallographic structure of Hofmann-Endell-Wilm[74] is the most accepted model of montmorillonite, which corresponds to a phyllosilicate (smectite group) with the general constitution: (Al1.67, Mg0.33)2Si4O10(OH)2 · nH2O[72,75]. Na+, K+, Ca2+, and other cations such as Co2+, Ni2+ and Cu2+ easily complex with montmorillonite. When occupied with Ca2+, the pH value of the T-T interlayer space is 7.5. In the interlamellar TOT-TOT space a big amount of water, hydrated metal cations and organic compounds can accumulate. Like silica, alumina and carbonaceous materials, clays show a striking ability for retention of dangerous or toxic compounds and are used as adsorbent agents for decontamination of plaguicides, pesticides, herbicides, etc.[76].
In addition to simple anions and cations[77], numerous organic compounds such as alcohols, amines, ethers, ketones, urea, phenols, glycols and polyglycols, benzene, benzoic acid, pyridine, anilines, soaps, surfactants, etc., can easily insert within interlamellar spaces of the silicate sheet from montmorillonite forming interlayer inclusion complexes[77-79]. Likewise, interactions of montmorillonite with several polymers (polypeptides, polysaccharides) has been described[56,80].
Intercalation of homo and heterocyclic compound into montmorillonite occurs with simultaneous stacking at low concentrations[81]. The adsorption of purines, pyrimidines, and nucleosides by montmorillonite has been early studied[82,83], but no structural aspects of their interactions have been investigated. Thymine and uracil were not adsorbed alone and the presence of adenine was necessary for appreciable adsorption, likely due to H bond formation with the clay T sheets[84].
The capacity of montmorillonite to induce dye aggregation is also well known, the stacking process being detected by spectroscopic methods[37,38,85-88]. Most of the inserted compound bind to T surfaces of clay by electrostatic and dipolar forces, as well as by H-bonding and coordination to metal cations[6]. Although lacking mechanistic details, montmorillonite and other clays have been considered possible scaffolding for the prebiotic formation of polynucleotides[89-91]. Likewise, the hydrolysis rate of phosphate-diesters could be prevented due to their inclusion within the montmorillonite matrix.
6. The Molecular-Engineering Zipper Model
A self-assembly process based on the stacking of nucleosides along the silicate surfaces of montmorillonite is the key feature of the clay-induced antiparallel stacking. As the polymeric chain with inverted stacking appears like a molecular zipper (Figure 6 and Figure 7), the proposed mechanism for the prebiotic origin of RNA can be named a "zipper" model, mainly regarding the two steps inside the montmorillonite cage. Interestingly, a process of reversible NAs denaturation and reassociation has been also named a zipper mechanism[29]. During reassociation, nucleation occurs by stacking new base pairs over the three first ones, allowing the double-helix formation by a cooperative zipper closing process.
A more realistic model of the resulting ladder-like auto-intercalated and zipper structure is here shown in atomic detail (Figure 8), and it is a key intermediate in the prebiotic formation of RNA molecules. The model has no unreasonable stereochemical contacts. Both stacking of continuous or alternate purine and pyrimidine nucleosides results in regular or irregular (aperiodic) but non-coding base sequences.
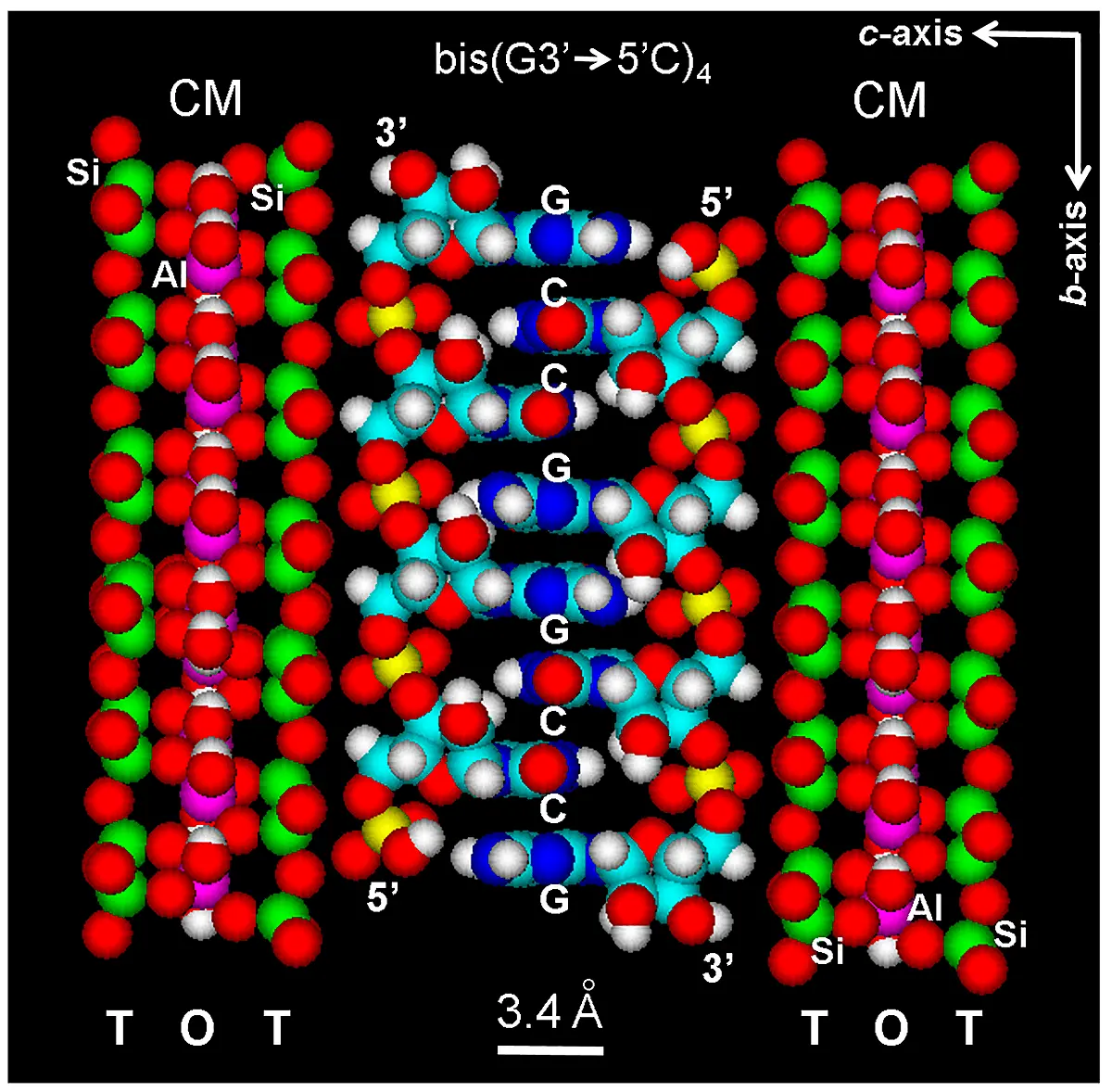
Figure 8. Space-filling (atomic volume) zipper model of the auto-intercalated and antiparallel bis(G3'→5'C)4 tetramer assembled within the interlayer space of the microcrystalline three-layered montmorillonite clay matrix (CM, according to the model of Hofmann-Endell-Wilm[74]), with sheets TOT-TOT (silicate tetrahedra: T, alumina octahedra: O). Crystallographic axis are indicated (top, right). Si: green; Al: violet. Color code of elements is C: cyan, O: red, N: blue, H: white, P: yellow.
Regarding auto-intercalation, a ss-model for auto-intercalated parallel poly(dC) was early proposed[92]. It was also suggested that models for base-mismatched antiparallel oligo(dA) could form either bulged-out ss-loops or auto-intercalated antiparallel chains[92]. In the case of the telomeric sequence (G4T2) ∙ (A2C4), it is tempting to speculate that, in addition to the (C, A) hairpin[93], the involved CCCCAA and GGGGTT sequences can also form two independent auto-intercalated structures.
It should be noted that ss-segments form random-coils (e.g., some regions of rRNA, tRNA and mRNA), which are stacked arrays of purine and pyrimidine bases surrounded by a helical ribose-phosphate ss-backbone. This linear structure has been widely confirmed[26,29,30]. In the transition stage between ss- and ds-RNA of the zipper model, the appearance of the ribose-phosphate chain as an unwound ladder-like structure is similar to that of both DNA fully intercalated with[(bipy)Pt(en)]2+[94], and unwound DNA[95].
Taking into account the spectral changes related to aggregation of aromatic chromophores[37,38], a spectroscopic study of the changes induced by montmorillonite as model substrate for stacking of guanosine has been performed. The absorption spectrum of guanosine shows a hypsochromic shift (from 252 to 247 nm) as well as a strong hypochromic effect in the presence of montmorillonite (Figure 9), a fact in complete agreement with other studies[46,49]. As expected, a striking hypoemission with red shift (from 350 to 360 nm) is also observed in the fluorescence spectra of solutions of guanosine mixed with montmorillonite. According to accepted interpretations, hypochromism and hypoemission of NAs is attributed to base stacking[30,49,96]. These spectral changes clearly indicate that this clay induces a strong aggregating effect on guanosine, which remains inserted as stacked dimers into the interlamellar spaces between silicate layers.
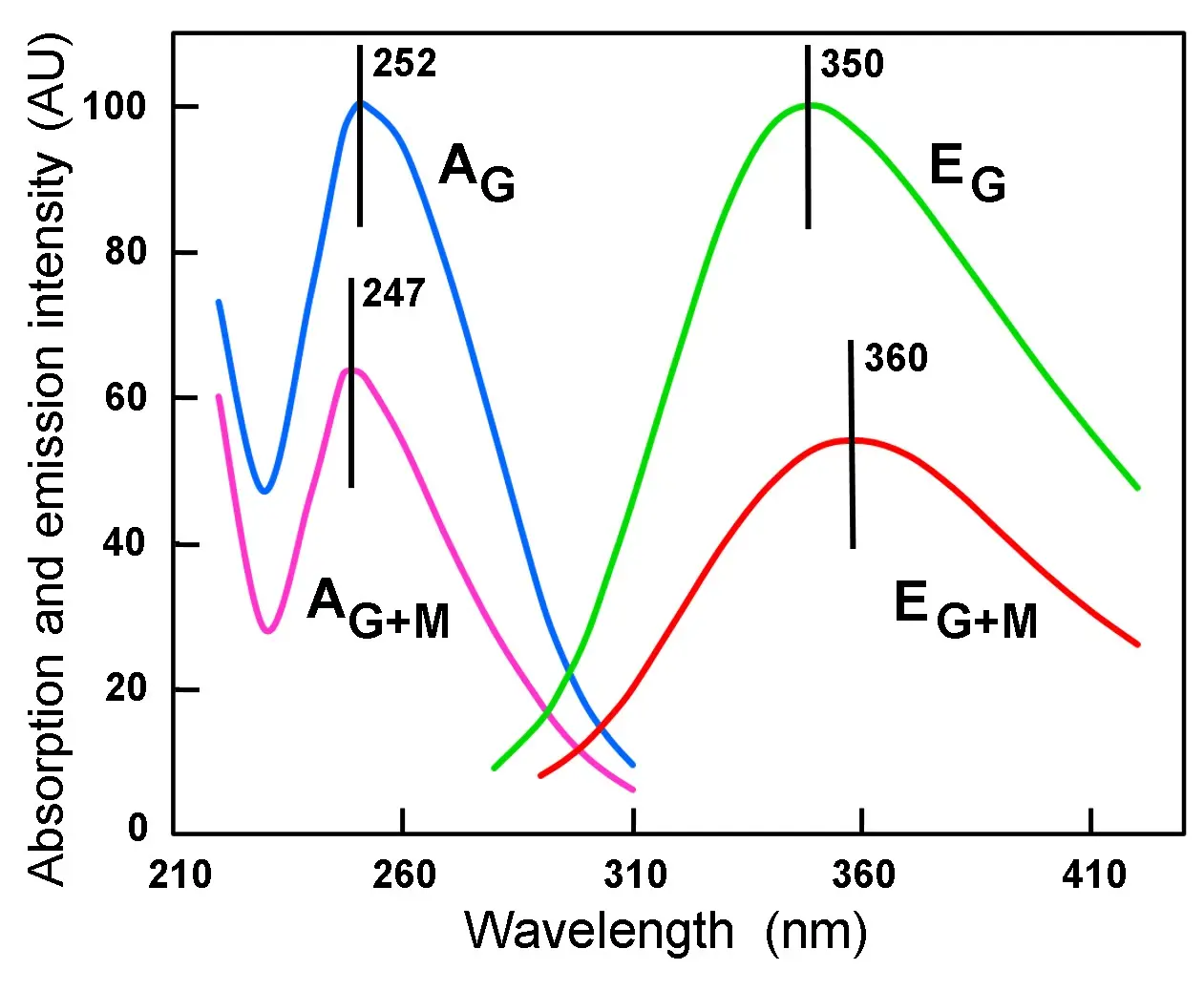
Figure 9. Normalized absorption (A) and emission (E) spectra of 9 x 10-5 M guanosine in distilled water, either alone (G) or in the presence of 0.2 mg/mL montmorillonite (G + M), showing the hypochromism and hypoemission induced by the clay. The position of A and E peaks (in nm) are indicated. Excitation wavelength: 260 nm.
Regarding fluorescence of NAs, it is known that 260 nm-excitation induces near UV emission, which is due to purine and pyrimidine bases[97,98]. Likewise, unstacking of bases after NAs denaturation causes hyperchromism with blue shift and hyperemission with red shift[26,29,46].
MOs studies on the modeled structure shown in Figure 8 reveal that most energy states in stacked guanosine have partially fused LUMOs at low energy levels. At higher levels, fusion is more general and expands to the column of auto-intercalated bases, which confirms that base stacking within the clay is responsible for the spectral changes of guanosine. An example of this phenomenon is the extended fusion pattern that occurs in the case of LUMO+9 from the auto-intercalated bis(G3'→5'C) 4 (Figure 10), which shows massively coupled π-electron interactions.
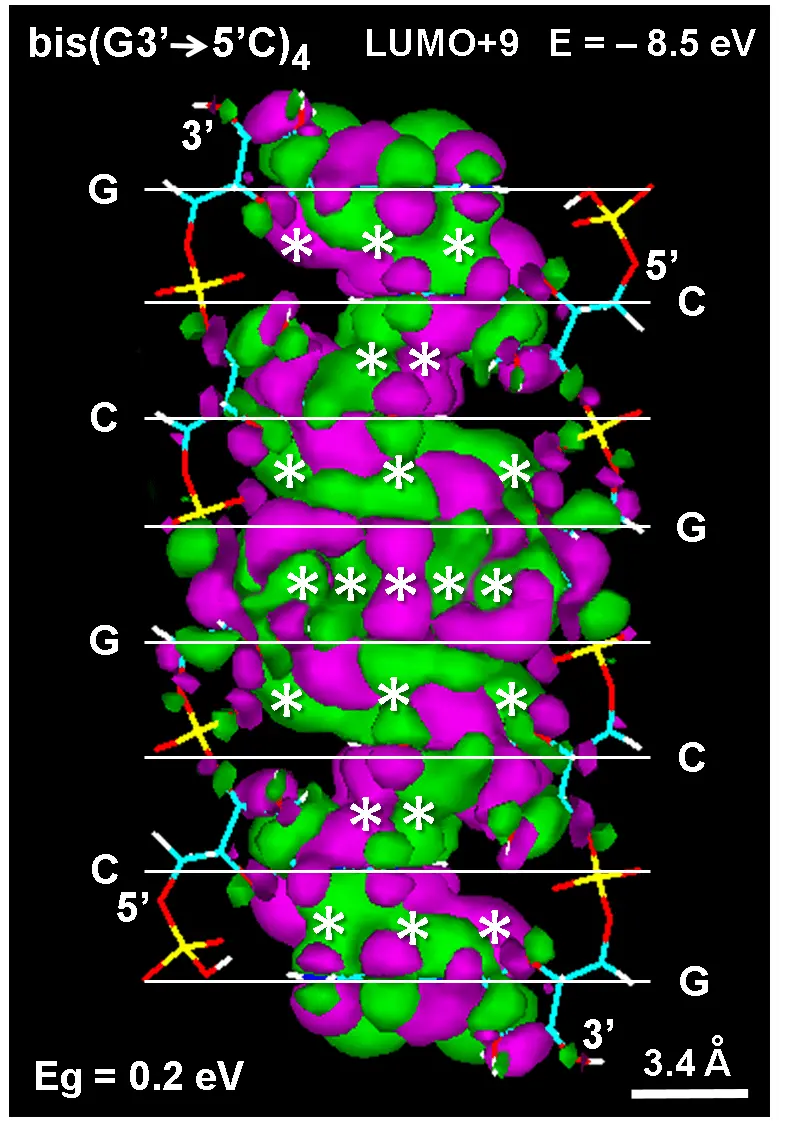
Figure 10. (A) Lateral view of the auto-intercalated and antiparallel bis(GC)4 model showing the electronic structure of the LUMO+9 energy state at -8.5 eV. Hückel method (788 orbitals, Gouraud shaded 3D isosurface, orbital contour: 0.0001). MO lobes from π* electrons have different colors. Asterisks denote massively fused LUMOs. O3' and O5' ends are indicated. Horizontal white lines show the position of guanine (G) and cytosine (C). (B) Distribution of HOMO (VB, blue) and LUMO (CB, red) energy values. Observe the small energy gap (Eg = 0.2 eV) between the valence and conduction bands. LUMO: lowest unoccupied; HOMO: highest occupied molecular orbital.
Interestingly, MOs of intercalated dyes between aromatic layers of graphitic materials (e.g., graphene, polydopamine, graphite-like eumelanin models, etc.), appear also fused due to the coupling between their π-electrons and those of the graphitic matrix[39-41]. This effect is just the observed here along stacked guanine bases of an auto-intercalated antiparallel ss-oligonucleotide model.
After released from the clay sheets under appropriate conditions of water temperature and pressure, osmotic swelling, pH and salinity[77,99], auto-intercalated polynucleotides lose the stacking between bases from different chains and become free ss-chains with a random coil conformation. A considerable winding and shortening of the ribose-PO4 chain due to new requirements for base stacking in the same chain (to reach from 6.8 to 3.4 Å) must occur by inclination of ~ 50-60º from the vertical axes and by changing the bond angles. These geometrical changes just result in a necessary helical conformation of the two antiparallel and WC-paired A-type ds-RNA that can be seen in Figure 11.
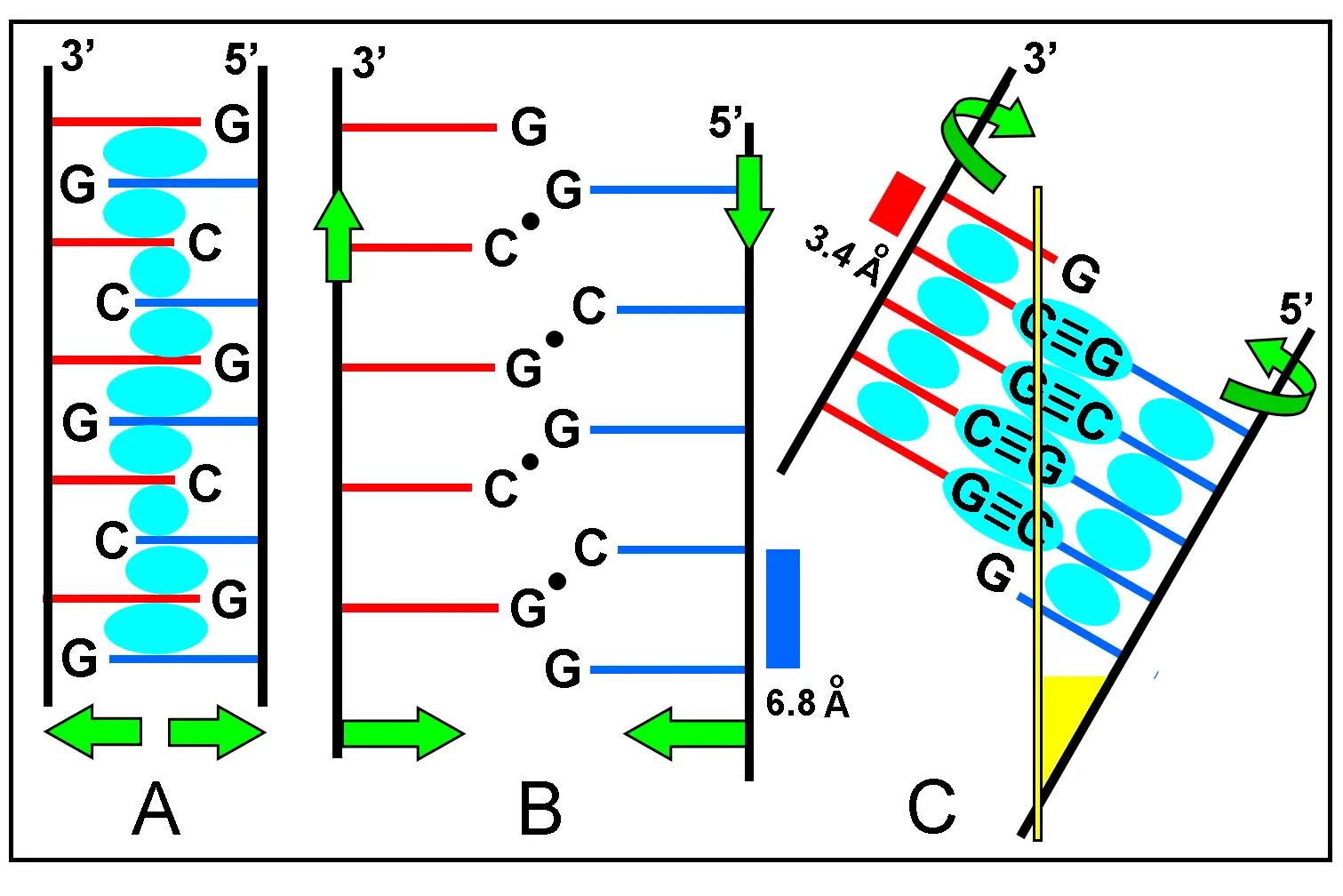
Figure 11. Schematic view of ds-RNA formation. (A) Initial step of the molecular zipper model, showing chains (black lines) with auto-intercalated nucleotides. Bases from different chains have red and blue colors. Green arrows indicate chains separation after detachment from the clay matrix; (B) Dissociated and unstacked polynucleotides form a random coil and search for WC base pairing (black circles) by linear displacement and shortening of chains (green arrows). This structure has yet an unwound configuration; (C) After pairing and stacking again, the two polynucleotides form the helical and antiparallel ds-RNA structure. Green curved arrows indicate winding of chains to reach adequate base pairing. Observe the inclination of the chain respect of the vertical yellow axis. The inclination angle corresponds to the yellow triangle at bottom. The cyan areas represent overlapped MOs from stacked bases in (A) and (C), resulting more stabilization of the RNA duplex by H-bonding. WC: Watson-Crick; Mos: molecular orbitals.
The average angle between the glycosidic N-C1' bonds from adjacent bases in the original unwound zipper model is about 22º, and the WC-edge angle for base pairing in the auto-intercalated nucleotides is 55º (Figure 4). Therefore, the final formation of a helical ds-RNA requires a rewinding of 33º (55 - 22 = 33) in the two chains, to reduce the WC-edge angle to 0º.
The molecular-engineering zipper model is based on preferential hydrophobic interactions and self-assembly of nucleosides within a montmorillonite cage, and can be viewed as sequential stages of different processes (Figure 12). It must be noted that the dominant forces of interaction between nucleosides comes from the stacking of their constituent bases[29,50]. The first and second stages (A and B) of the model give the zipper name to the ordered ladder-like, auto-intercalated ss-oligonucleotide, with montmorillonite serving as cage or scaffolding for the assembly of building units. As expected for ss-RNA, the corresponding ss-chains (stage C) have a considerable freedom degree and can adopt variable random coil conformations[29]. It is known that ss-chains of poly(A) and poly(C) are helical structures[29].
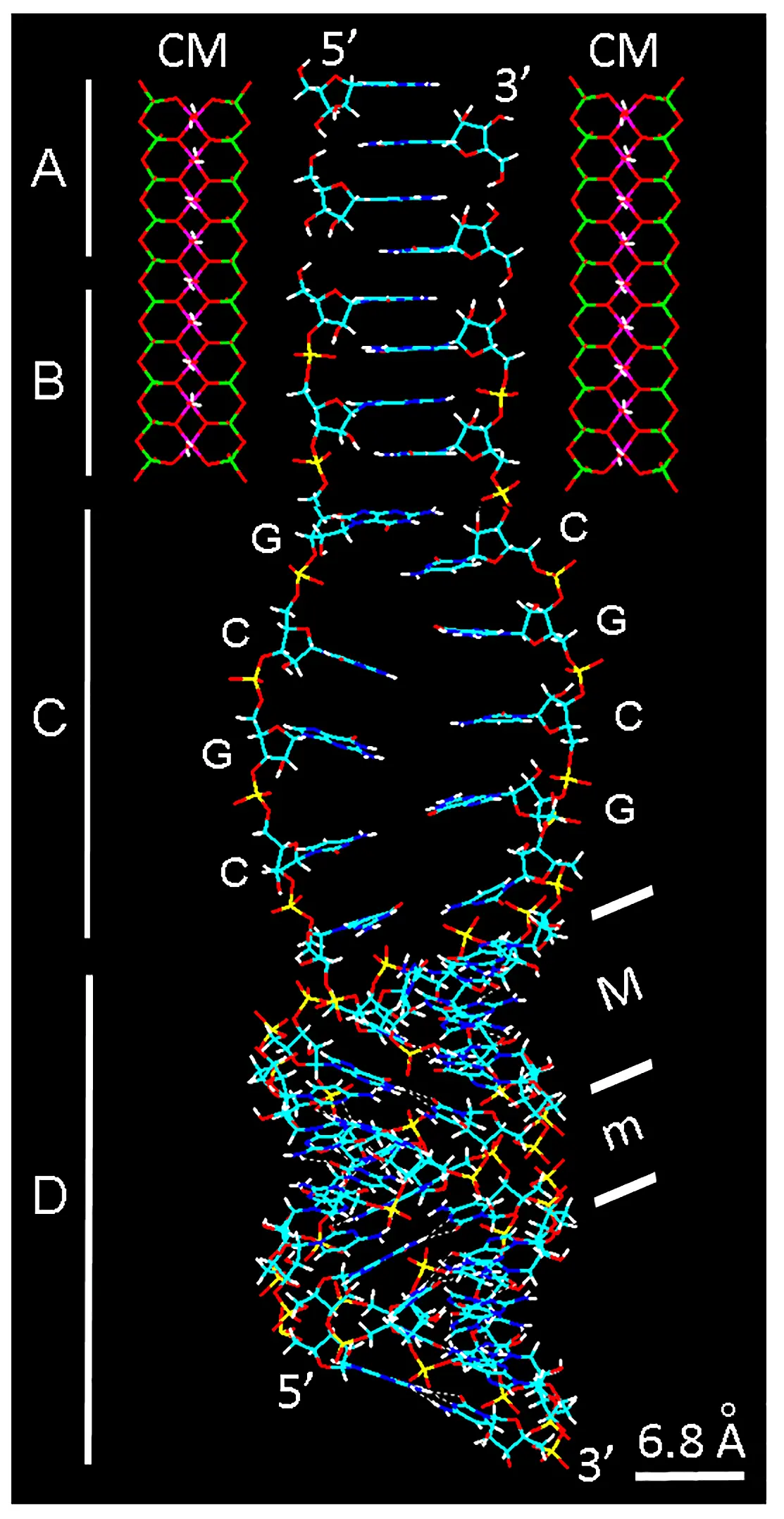
Figure 12. Graphical abstract of the molecular-engineering zipper model showing the sequential self-assembly of the auto-intercalated ss-RNA within a montmorillonite CM, followed by formation of ds-RNA. The wire model represents the four key processes of RNA building represented as vertical segments on the left. (A) Antiparallel stacking of nucleoside dimers; (B) Phosphate bridges forming 3'-5' ribonucleotide chains, after reaction with high-energy triphosphates; (C) Dissociation of ss-RNA chains (as random coils) from the matrix, with G and C indicating guanine and cytosine bases; (D) Formation of the right-hand helical A-RNA duplex from the complementary base pairing in previous random coils. Inclined bars indicate the crests of ribose-phosphate chains and delimit the minor (m) and major (M) grooves. Note the tilt angle of base pairs (~ 13º) and ~ 65º inclination of the helical chains in relation to the vertical axis. Color code of elements is C: cyan, O: red, N: blue, H: white, P: yellow. CM: clay matrix.
Once formed the antiparallel ss-chains, the origin of ds-RNA helices is a conceivable consequence, in dependence on the base pairing complementarity and the shortening-winding process of chains. It is worth to note that NAs helicity just allows the helical transitions between all the other helical NAs: right-hand A↔B DNA types and left-hand Z form. Likewise, a Z-RNA form has been described[100,101].
Irregular base sequences along the auto-intercalated intermediate (e.g., consecutive 1 to n dimers with the same bases) will produce unpaired terminal RNA segments and internal bulge loops. Free ends with unpaired bases can then close by PO4 bridges retaining the 3'-5' orientation and forming a loop structure.
On the other hand, several variations related with the general NA architecture could also be possible. AT sequences in some hairpins from DNA duplexes with W-C H-bonded parallel strands instead of antiparallel ones have been found[102], which does not invalid the acceptance of the general antiparallelism of NA duplexes.
7. Origin and Replication of RNA Structures
Along the formation of ss- and ds-RNA, numerous simple and complex structures and conformations with biological significance are possible. Following the zipper model, the original and simplest oligomeric unit would be a ds-RNA (stem) with paired and unpaired open (free) ends. These ends can be closed by a phosphate bridge, giving hairpins such as the typical stem-loop motif (Figure 13), which relates the zipper model with the biochemistry and genetics of typical RNA structures (e.g., tRNAs, infective agents such as viroids and virusoids, etc.).
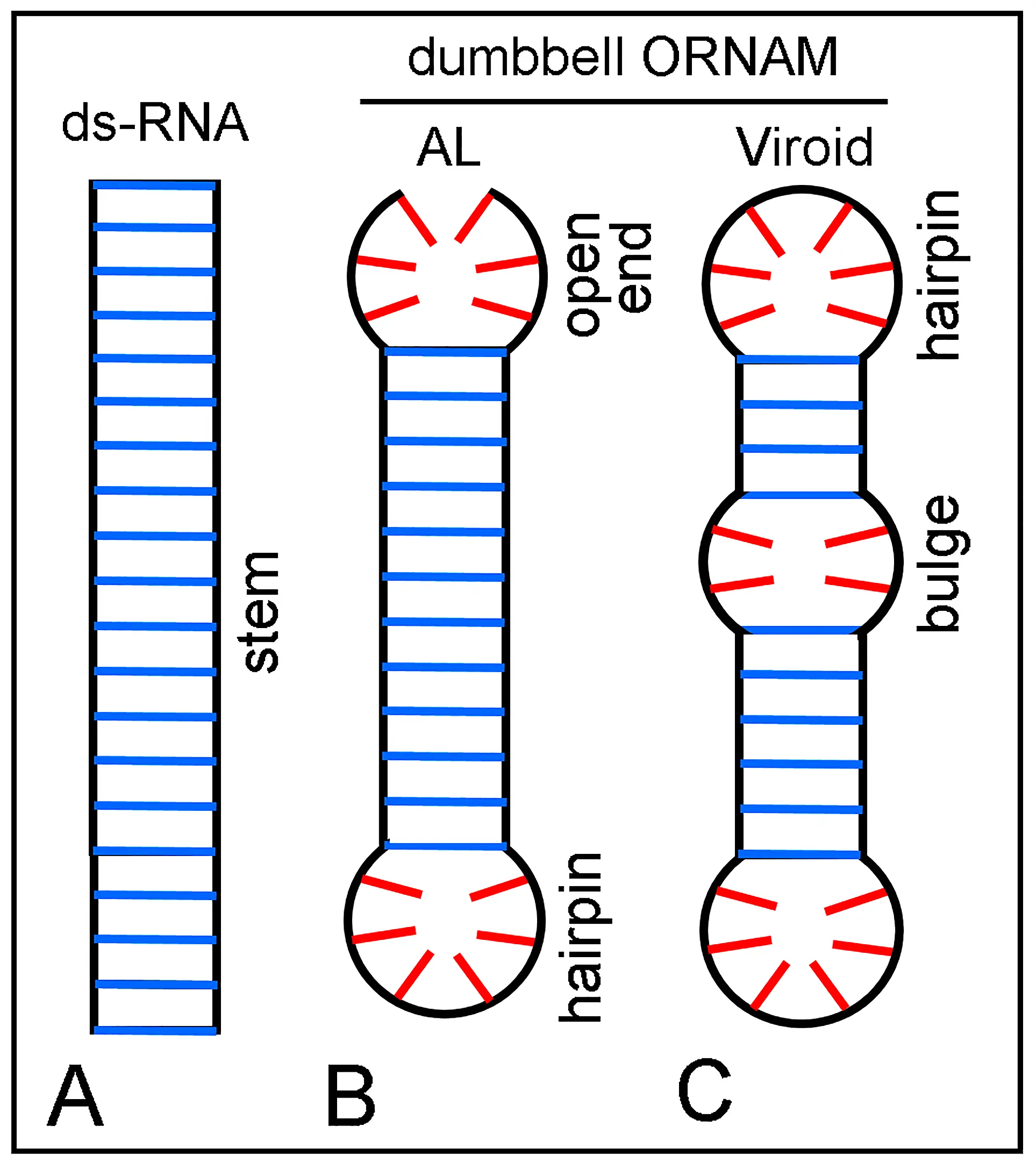
Figure 13. Schematic representation of stem ds-RNA segments with terminal and internal unpaired loops. Vertical black lines are ribose-phosphate chains. Blue and red lines indicate paired and unpaired bases, respectively. (A) Stem of a wholly paired ds-RNA; (B) Stem with a closed end (hairpin loop) and an open (unpaired) end, which correspond to an AL sequence (see text)[103]; (C) Stem with hairpin loops and internal unpaired loops (bulge). B and C correspond to the basic structure of an ORNAM, namely dumbbells and rod-shaped viroids and virusoids. AL: Archetypal loop; ORNAM: Original RNA Motif.
It is known that short ligation-activated oligonucleotides (i.e., tetra- to dodeca-nucleotides) undergo efficient intramolecular ligation (cyclization)[18,104], closing the free unpaired ends of ss-chains to form dumbbells. Using suitable nucleoside dimers and agreeing sequences, the resulting two ss-RNA stem chains are expected to form a well-paired RNA duplex. However, with some unsuitable bases, small or large non-complementary ss-chains will be formed, remaining as intercalated pinholes, bulges or terminal unclosed loops. This is just what occurs in the case of viroids and virusoids, but this does not signify that they are directly generated through the zipper model.
It is worth to note that these stem-loop motifs are also present in all natural (biological) RNAs, and their size, position, specific sequence and catalytic properties are characteristic markers for different RNA types. A few examples are the highly complex secondary and tertiary structure of typically "L" folded tRNAs, 16S/23S (Bacteria) and 18S/28S (Eukarya) rRNAs, 5S rRNA, small nuclear RNAs (e.g., U1 RNA, etc.), RNase P RNAs, small interfering and non-coding RNAs, micro RNAs, tiny noncoding RNAs, mRNA regions, etc. They contain stems of minihelices, hairpin loops, internal loops, bulge loops, hammerheads and double hammerheads, cloverleaves, junctions, pseudoknots, etc.[26,29,30,105-114]. At present, it is outstanding that also DNA can form microstructures named DNAzymes with catalytic properties like ribozymes. An example is the 10-23 DNAzyme catalyst (a short DNA fragment with a loop between two arms) that break specific regions of ss-RNA[115,116].
The comparison between the possible ORNAM products formed according the zipper model and the simplest RNAs such as viroids, virusoids and AL sequence is amazing. In fact, there are striking structural similarities between all these RNA motifs. Figure 13B,C should be compared with both the rod PSTVd viroid[117], and the proposed AL (Archetypal Loop) sequence of 22 nucleotides from the archaeal 5S tRNAs of Rhodobacter sphaeroides : UGAAUGGUA/C/UGCCAUUCA/ AGA[103].
In the last few years, an increasing interest was given to the structure and properties of small self-paired circular RNAs or similar motifs. An example is the fifty-five base hairpin fragment from the R17 virus[118]. Self-paired rings (in the form of dumbbells) have received special attention[67], because they are highly stable compact structures that are resistant to degradation by nucleases. In keeping with this, the concept of "ring world" has been recently introduced to incorporate the organization and function of small circular NAs (AL elements, viroids, virusoids, etc.)[119]. Interestingly, the already described ORNAM structure also belongs to the ring world.
Viroids were firstly described by Diener in 1971[120], and their history, composition, properties, and pathogenicity have been reviewed[12,117,121-123]. Virusoids are similar to viroids in respect to size and circularity, but do replicate only as part of a larger plant virus. Viroids are independently replicating, covalently closed circles of small ss-RNA ranging from 246 to 375 nucleotides, and do not contain capside, proteins nor lipids. In spite of their small size, they are very pathogenic to many plant species.
These simple RNA molecules are either rod-like (stem) or branched (hammerhead) structures showing base-paired regions interspersed with ss-hairpin and ss-bulge loops[124]. Viroids do not code for any protein and their replication mechanism uses RNA polymerase II, a host cell enzyme normally associated with synthesis of mRNA from DNA. A rolling circle model has been proposed for replication using the viroid RNA as a template model[125]. Viroids are often ribozymes, having catalytic properties that allow self-cleavage and ligation. The activity of suitable functional ribozymes in folded sequences (hammerheads) occurs in longer RNA structures such as virusoids[109,112]. Note that the majority of modern ribozymes catalyses phosphoryl transfer reactions, either hydrolysis or transesterification[126]. Diener's hypothesis[127] proposed that the unique properties of viroids could possibly make them "living relics" of an ancient, catalytical and pre-cellular RNA world. This interesting view was mostly forgotten until 2014, when it was resurrected by Flores et al.[15], and now this conception is currently accepted.
In relation to the prebiotic RNA origin, the likely formation of a random non-coding ss-RNA chain has been already described in the Section 4. However, replication of a coding ss-RNA on a template is a different problem. Regarding possible pathways of prebiotic RNA replication in a non-enzymatic scenario, a rolling circle replication of viroids has been proposed[125], but it is not viewed at present as a viable model. The use of adenosine-5'-phosphorimidazolide as reactive precursor has been suggested[66]. NaCl-montmorillonite catalyzes RNA synthesis from activated imidazolide-nucleotides and facilitates homochiral selection[89,91] for spontaneous untemplated and templated synthesis of RNA. A virtual circular genome model (VCG, a collection of short circular 12-nucleotides) has recently been suggested for prebiotically plausible nonenzymatic RNA replication cycles during the origin of life[128,129].
The use of activated dinucleotide precursors with a reactive group (e.g., 2-amino- and 2-methyl-imidazolium) has now proved very adequate to recognize the template by two base pairs[68,130]. Furthermore, highly reactive imidazolium-bridged, activated RNA trimer helpers greatly accelerate nonenzymatic primer extensions, because of the structural preorganization afforded by an extended helical geometry[69]. Nucleoside-5'-phosphoryl-imidazole reacts with remarkable efficiency on a suitable ss-template to give activated precursors and oligomers in aqueous solution[89,91]. The prebiotic occurrence of imidazole derivatives is not implausible[66]. Chemically activated short oligonucleotides also cyclize efficiently using N-cyano-imidazole[18]. It is worth noting that as aminoacyl-imidazoles give peptides in aqueous solution[131], the imidazole group (e.g., the peptide-forming reagent, N,N'-carbonyl-diimidazole) may provide a link between prebiotic and biotic reactions for protein synthesis.
The simplest RNA molecules formed according to the zipper model (e.g., the ORNAM dumbbell), as well as AL, VCG, viroids and virusoids (Figure 13) could originally replicate from available precursors self-assembled in both a stem-loop junction and ads-RNA stem. A graphical model for the possible non-enzymatic RNA replication can be seen in Figure 14A. The possible formation of an RNA tetraplex segment is also shown, which results from a very simple replication mechanism, leading to a compact replicated RNA dumbbell product, and relaxing the turn stress generated during replication unwinding.
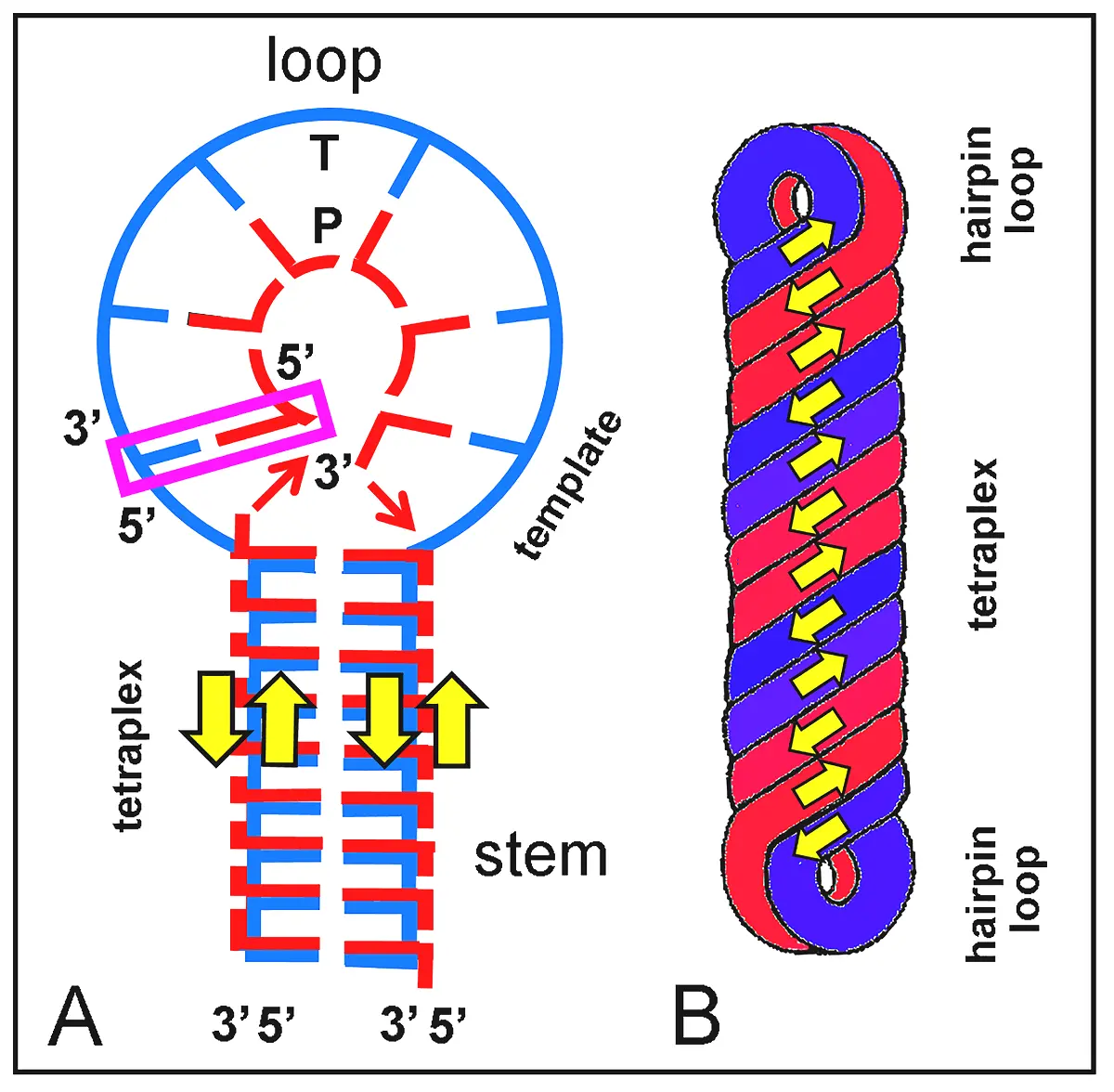
Figure 14. Replicative stem-loop model for an RNA structure (e.g., ORNAM). Replication proceeds by polymerization of paired activated nucleotides or phosphorylation of also paired nucleosides as a primer (P, red) on an ss-RNA template (T, blue). When no primer exists, the first nucleotide on the newly synthesized chain serves as initiator. Color bars are bases from identical chains. (A) Schematic representation of replication taking place at the stem-loop junction (replication initiator, violet rectangle) and progressing to form a duplex loop region. Precursor nucleotides and their components (here non-proportional in size) are L-shaped units (Figure 6). In the stem, an RNA tetraplex is formed by self-assembly of precursors following the "tetris" replication model. (B) Simplified morphology of the replicated dumbbell RNA according to the process shown in (A). The replicated ORNAM appears as a helical rod structure with 3 right-handed turns along the stem tetraplex and very short hairpin loop duplexes. Yellow arrows point out the antiparallel 3' → 5' orientation of ss-chains.
In the stem-loop replication model, growing of ss-polynucleotides may occur by direct pairing of nucleotide-triphosphates or by phosphorylation of nucleosides previously paired with the template, either using polyphosphates (Figure 7), imidazolium or other energy-rich catalysts. Both stem-loop junctions from a dumbbell RNA could initiate replication simultaneously, and a relaxed replicated form should appear as a ds-circular chain. However, when observing this possible replication mechanism, it is logical to speculate that a tetra-stranded stem could also occur as a natural consequence of the replication process.
In addition to the self-assembly of complementary precursor on a small ss-loop from the ORNAM structure, it is exciting to consider that an RNA tetraplex could also represent an interesting and possible alternative mechanism for conservative replication of RNA. This novel process would be based in the self-assembly of activated nucleotides or nucleosides in the presence of high-energy phosphates, which could form complementary H-bonds along the major groove of RNA duplexes. The self-assembly of RNA precursors can be graphically described as the falling or docking of "tetraminoes" pieces in the Tetris game. In keeping with this, the "tetris" model proposed here for prebiotic RNA replication is based on the occurrence of available and complementary surface targets on the major groove of ds-RNA stems (Figure 2A and Figure 13A). Advantages of this simplistic model are that it does not require previous denaturation of the parental RNA duplex, and duplex segregation from a tetraplex involve simple uncoiling.
Multiplex NAs structures are worth to note. Three-stranded helical NAs are well known[26,111,132-134] and their structure and biological functions have been recently reviewed[135-137]. Base triads are found in tRNA, synthetic ribo- and deoxyribo polynucleotides and specific DNA regions[29,111], the second (extra) pyrimidine chain binds to purines in the deep major groove and runs parallel to the purine chain[30,93]. Conformational forms of NAs are widely known[26,28,29,39,138], as well as specific and complementary H-bonding between paired bases[29,139]. However, some dynamic aspects of NAs should be considered, e.g., unwinding and winding in a semi-conservative replication process. During separation of RNA strands, every turn of the original duplex must to be unwound so that the strands can be dissociated, each of them becoming finally one strand of the replicated duplexes.
In a prebiotic condition lacking gyrases and topoisomerases, unwound or left-handed RNA duplexes could accommodate the torsional stress generated by replication. Relaxation of the wound ds-chains could take place by the free rotation of linkages at the ss-loop or circular chains. From the topological standpoint, the central ds-stem regions should also accumulate unwound turns that could then result in a right-handed intercoiled ds-RNA during rewinding with possible H-pairing of two duplexes along their major grooves. This exciting solution would allow the formation of a transient right-handed RNA tetraplex in the final stage of replication (Figure 14A,B).
Obviously, the non-enzymatic copying of ds-RNA results in two duplexes that must be dissociated to allow the next replication round. Thermal denaturation, pressure and pH fluctuations, solvent viscosity and other environmental changes can readily dissociate short RNA tetraplexes from stems or contained in hairpins, AL and ORNAM structures.
Regarding tetraplexes, several authors have suggested and found the occurrence of base tetrads (Figure 15) in theoretical and crystallographic studies on DNA[140-145]. A known example is the occurrence of G·G·G·G tetrads and tetraplexes (quadruplexes) (Figure 15A). In the cases of A·T-T·A tetrads (Figure 15B), a DNA tetraplex can be formed with about ten base tetrads per turn, with base planes normal to the helix axis and glycosidic bonds set at the corners of a rectangle with sides of 10.8 Å. Figure 15 B and Figure 16 illustrate two tetrads in order to form four-strand structures with a very solid core of stacked and H-paired base tetrads (side-by-side base pairs)[140,144], allowing the formation of helical tetraplexes[140,141,144,146-148], even lacking the homologous pairing in base tetrads[149,150]. Hairpin loops need contain only two base pairs to be closed[147].
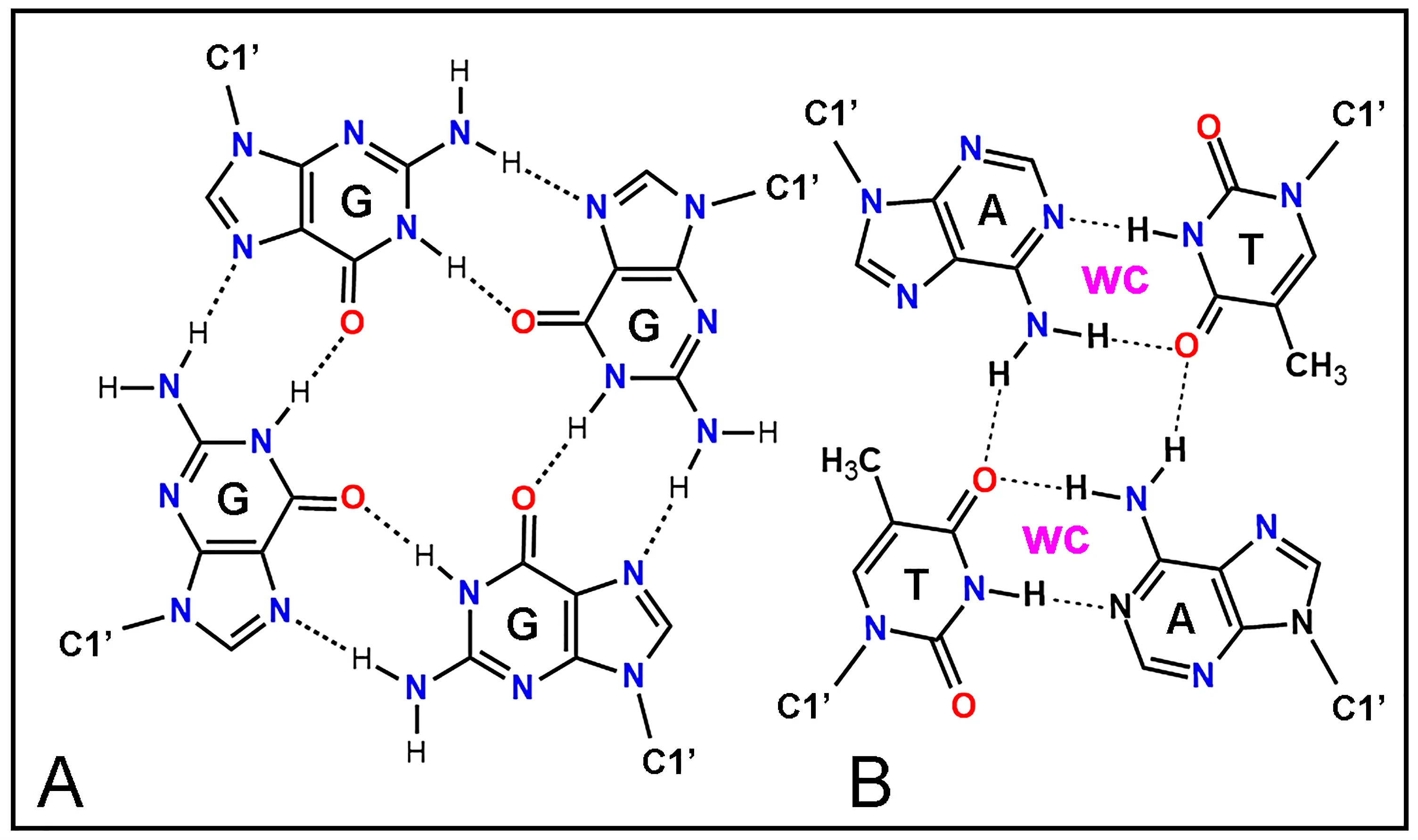
Figure 15. Chemical structure of base tetrads as building blocks for tetraplexes using H-bonding (dashed lines). (A) G·G·G·G tetrad. (B) A·T-T·A tetrad formed by two complementary A·T dyads with Watson-Crick (WC) base pairing. C1' indicates the glycosidic atom of pentoses that binds with N of bases. A: adenine; G: guanine; C: cytosine; T: thymine.
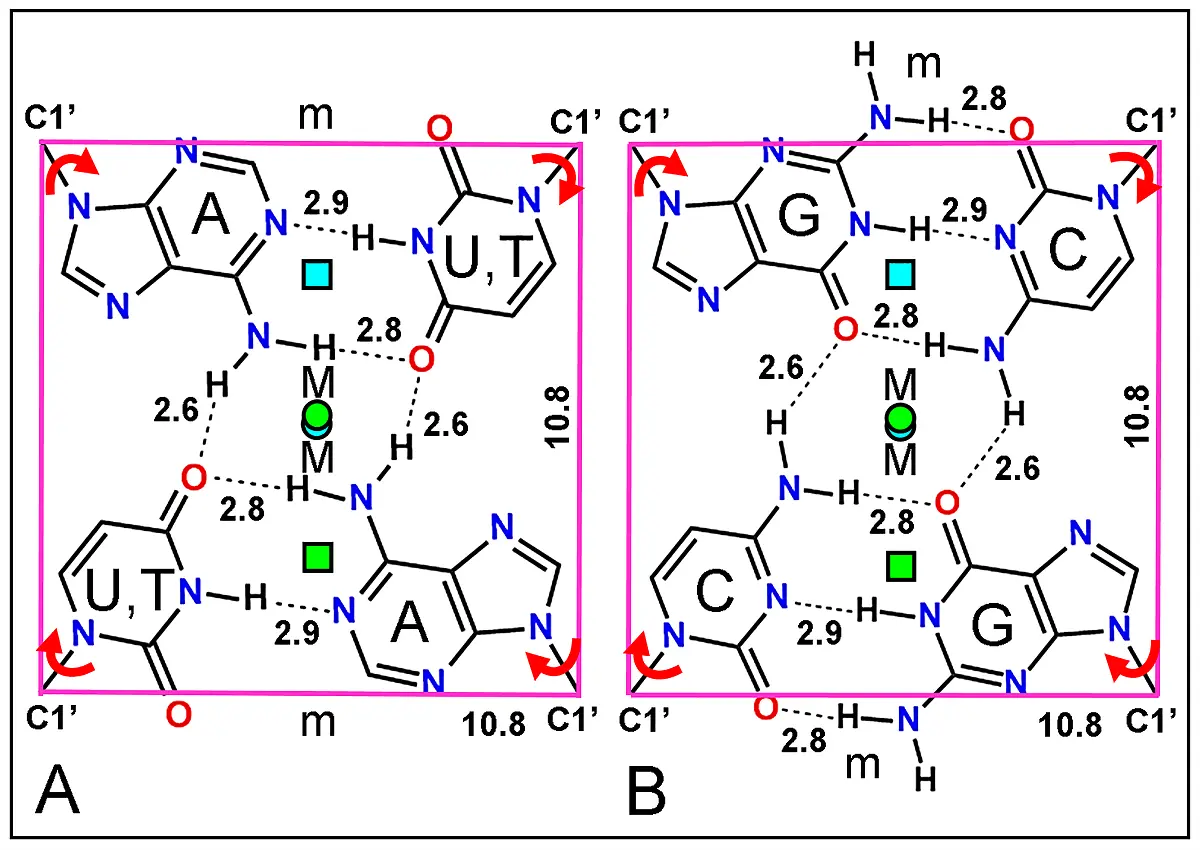
Figure 16. Structure of two complementary base dyads. (A, B) Symmetrical AU(T)·AU(T) and GC·GC base tetrads formed by two duplexes faced by the side of the major groove using the complementary lone electron donor from O and H-acceptor from N atoms code[139]. Each base tetrad appears inserted within the corners defined by the C1' atom of riboses. Numbers indicate H-bonds (dashed lines) and square length in Å. The two axis from B-DNA duplexes (cyan and green squares) do not coincide (for AT base pairs), whereas those from A-RNA are almost co-axial (cyan and green circles). Curved red arrows show the possible 180º rotation of glycosidic C1'-N bonds allowing orthogonal changes in the symmetric H-bonding pattern. Major (M) and minor (m) helical grooves are indicated.
Studies on complementary and symmetric base tetrads substantiate the formation of tetraplex structures invoked for DNA recombination[140,144,146,148,151-153]. DNA tetraplexes are also related with palindrome sequences[154,155], kinks and cruciforms structures[155,156], duplex-tetraplex Z- and B-DNA junctions[148,153], intercalated four-strand DNA[157] and other multi-stranded NAs[145,147,150,156].
Taking into account the possible DNA tetrads and two-duplex DNA tetraplexes[144-148], why not having a two-duplex RNA tetraplex as it was proposed before in this article? Obviously, the same tetrad geometry of deoxyribonucleotides could be shared by ribonucleotides (Figure 16).
In the case of RNA, the coaxial correspondence of both duplexes would be greatly better than for DNA. Note that in A-RNA, the helix axis is displaced far before the base pairs in the deep major groove, which provides more space to accommodate tetrads. Likewise, due to the 10º-13º tilt angle of base pairs in A-RNA duplexes, the tilt between duplexes in the tetrad increases to 19º. In keeping with this, the complementary duplexes retain their H-bonds pattern and improve the length between them from 2.6 to 2.8 Å. However, as occurs in the case of DNA[149,150], WC pairing between bases are not necessary to form RNA tetraplexes. By using a dumbbell structure, a simple pristine replication cycle can be formulated with loops and stem as template. After synthesis of the new RNA chains, the replicated ds-stem could form RNA tetraplex to release the torsional stress. Once replicated the dumbbell duplex, it can remain either as a regular and compact stem tetraplex (rod) or convert into elongated and circular duplex (ring), with formation of a left-handed supercoil. These would be reversible structures.
Most references to tetraplexes concern DNA, and it seems relevant that the RNA tetraplex here proposed could be similar to already described DNA tetraplexes. In order to facilitate the interpretation of the rather complex organization of these NAs structures, 3D images from both RNA duplex and RNA tetraplex were chosen to illustrate their main features. Accordingly, Figure 17 shows an RNA duplex and a stereochemically acceptable RNA tetraplex model, in which the four plectonemic, right-handed ribose-phosphate helices are clearly seen.
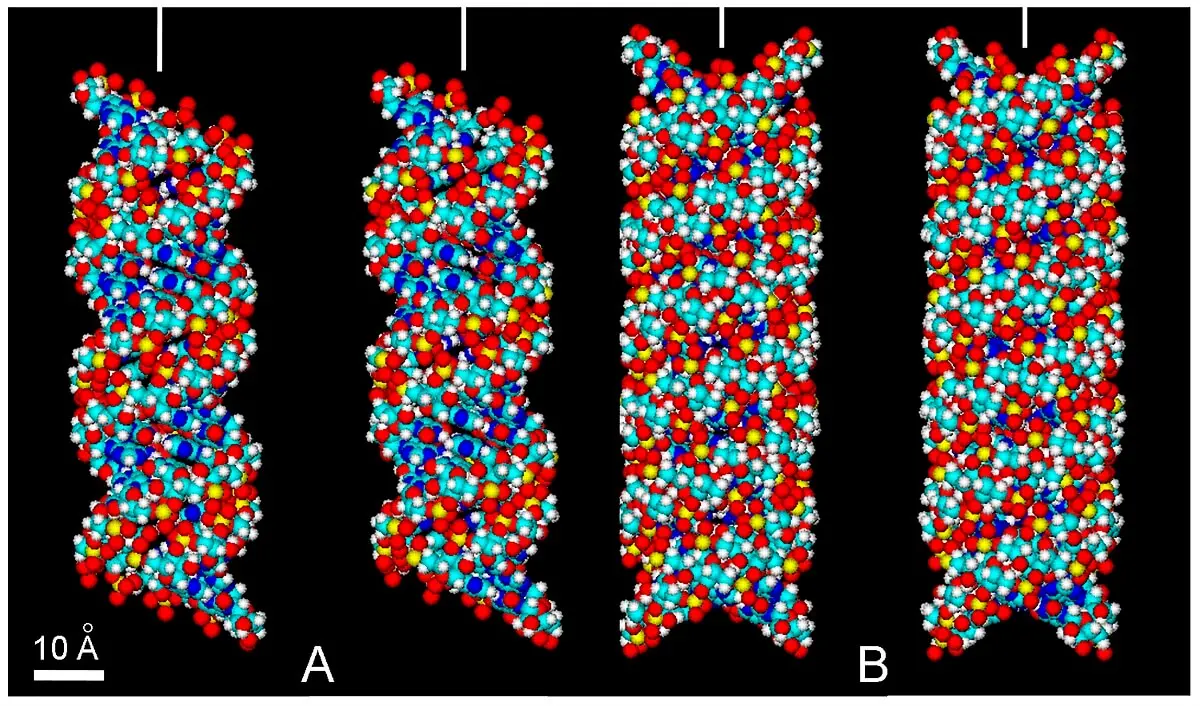
Figure 17. Stereopairs of atomic volume models of RNA duplex and tetraplex. (A) The RNA oligomer r(AU) 12 · r(AU) 12. (B) RNA tetraplex proposed here from two RNA duplexes shown in (A), using parameters described for poly (rG) quadruplexes[158,159] and DNA tetraplexes[141,152]. The RNA duplex and RNA tetraplex show different diameter (26 and 29 Å, respectively) and a turn length (92 and 96 Å, respectively). Major grooves of homologous duplexes are faced together, resulting in a very compact helical structure with narrower minor grooves. Observe that these duplex and tetraplex can be viewed as parental and replicated RNAs, respectively. In these RNA structures, there is a close approach of PO4 groups of neighboring chains across the minor groove (~ 4 Å), with metal cations bridging such narrow separation[160]. 3D view without optical aids can be achieved by relaxing eyes and focusing at infinity, followed by image fusion[43,161]. Stereopairs are easier observed by previous visual fusion of the vertical bars at top of the images, resulting in a single central bar. Color code of elements is C: cyan, O: red, N: blue, H: white, P: yellow.
Regarding topological aspects, it must be noted that for every turn of the WC helical duplex the helix is coiled once in the opposite direction[147,149,150,154]. There is a possible transition between tetraplex and supercoiled duplex forms. Taking into account the stem-loop junction replication model of a dumbbell ORNAM, it is tempting to propose that small circular ds-RNAs such as shown in Figure 13 and Figure 14B could represent original "living-like" molecular structures resulting from self-assembly and self-replicative events.
The occurrence of a possible A-RNA tetraplex proposed here seems to have been overlooked, probably because a clear biological role is still lacking. Torsional relaxation along the replication of closed circular RNA such as viroids and virusoids would be a possibility of using RNA tetraplex, either with paired or unpaired bases. It is noteworthy that one solution to the unwinding problem of DNA replication is the formation of a sequence-independent tetraplex, without pairing of the two base pairs. Tetrads and DNA tetraplexes of this type were found to be stereochemically possible[149,150]. Appropriate environmental conditions of temperature, solvation and ionic strength could induce the dissociation of two RNA duplexes from the parental tetraplex.
In addition, it is known that oligo and poly (rG), as well as poly (rI) easily form guanine and inosine tetrads and quadruple helices[29,158] (Figure 15B). A quadruple oligo (dG) was also proposed for the structure of chromosome telomeres, which contains tandem repeats of the sequence d(TTAGGG)[159,162]. Relations between G-quadruplexes and the origin of life have been quoted recently[163]. An "i-motif" from the telomeric repeat also corresponds to an auto-intercalated DNA tetraplex associated to the complementary G-quadruplex[164-166]. All these novel structures have a great interest for understanding the variable conformations and dynamics of NAs, which in turn are most probably related to their prebiotic origin.
8. Implications of the Zipper and Tetris RNA Models for the Origin of Life
Present studies on the origin of life involves multidisciplinary research mainly concerning geology, physics, chemistry and biology fields. More than half a century ago, early authors proposed that clay minerals might have played an important role in prebiotic synthesis[1,2]. More recently, interesting aspects of the RNA origin related to montmorillonite have been commented[59,89,90]. In the present work, a mechanistic rationale accounts for the self-assembling of RNA chains within a montmorillonite matrix, as well as for their prebiotic replication cycles. It can be assumed that pristine RNA structures were thus formed and developed in an RNA world[12,15], which is now considered the most plausible scenario for the origin of cellular life[5,6,13]. It must be emphasized that self-assembling is a well-known principle that govern the matter organization from atoms[167] to the molecular biosynthesis[19], including coenzymes and biopolymers[168,169].
The zipper model is based on self-assemblies and conformational changes of precursors, which finally result in a right-handed, helical and antiparallel RNA duplex. It is astonishing that this model stays in agreement with numerous molecular parameters, which allow explaining the RNA "phylogeny". Therefore, the correspondence between RNA structure and parameters such as geometric shapes, stereochemical fitting, free energy and bond lengths of precursors, etc., would be not a coincidence but a necessity for the origin of the RNA world (as well as for the life itself).
Obviously, the biochemical consequences of NAs structure (e.g., related to transcription and replication processes) cannot be argued to explain a priori the striking features of their helicity and antiparallelism. Likewise, it is worth to note that many nucleoside-like compounds (e.g., with glycosidic bonds between other types of sugars and heterocyclic rings) could also be available and then employed in the building of prebiotic biopolymers, but today, the unique sugar-PO4 -base polymers that remained are the usual RNA and DNA molecules.
Why purines and pyrimidines? These small planar, aromatic and functionalized heterocycles are particularly appropriate for self-assembly by stacking and for edge recognition and binding by H-bonding. At present, chemists have synthetized a big amount of heterocycles, but along the even prolonged prebiotic chemical evolution it seems logical to assume that only a smaller amount of natural compounds were available as building blocks, and purine and pyrimidine derivatives were possibly the most suitable.
Why ribose? Surely, numerous sugars formed in the prebiotic period by formose reactions. The possible use of threose or arabinose NAs (instead of ribose) has been reported[16,69]. However, only those that were rigid, cyclic and with hydroxyls placed at appropriate sites (such as β-D-ribose), could have advantages over other sugars for polymerization using a clay matrix. Thus, only the best-fitted heterocycles and sugars were conserved for building RNA polymers.
And finally, why phosphate? Several inorganic acids could be used as linking units between contiguous riboses. In addition to differences in chemical reactivity, they have distinct shape and size. Geometry optimization and energy minimization of possible ester bridges points out that bond length (in Å) between neighbor O atoms of several anions, are the following: nitrate-, 2.14; carbonate2-, 2.23; borate-, 2.34; sulfate2-, 2.56; phosphate3-, 2.71, and arseniate3-, 3.03. This indicates that a phosphate link (2.71 Å) fits better with the O3'-O5' distance between riboses (2.70 Å) than any other anion (Figure 7).
In agreement with the current conceptions in the RNA world, it is clear that the origin of RNA itself was an important event, involving the appearance of catalytic activity, self-replication, evolution[170-172] and finally integration in more complex prebiotic and then living organisms. Interestingly, ancestral hammerhead RNAs could evolve to more sophisticated ribozymes[109,118], as well as tRNAs and rRNAs from catalytic regions of proto-ribosomes, leading to the formation of peptide bonds and linking the pristine RNA world with the present DNA and protein world[173,174]. In a previous evolutive step, the zipper model explains nicely the origin of ss- and ds-RNAs, as well as the helicity and antiparallelism of NAs duplexes. In fact, it would be conceivable that under appropriate physico-chemical conditions and prolonged time, the molecular self-assembly of RNA polymers is practically inevitable.
It is worth to note that at present time, a highly complex semi-conservative mechanism has been demonstrated to occur for DNA replication, with segregation of the parental strands and formation of hybrid duplexes[26,30,175,176], Previously, a conservative mechanism involving a four-stranded DNA structure had been formulated, with parental strands remaining bound together and forming an entirely new duplex[177-178]. A similar but more complex DNA replication mechanism has been suggested by Bloch[179], based on the 180º-rotation of bases toward the major groove followed by WC pairing with incoming precursors. Likewise, Fong[180] and Löwdin[181] have proposed that a "replication plane" containing a tetrad of four complementary bases is energetically more favorable and has higher pairing fidelity than a replication plane with only two bases.
Just like it is today in the case of highly complex DNA replication, RNA transcription and protein biosynthesis, it would be expected that a prebiotic (non-enzymatic) RNA replication could be a simpler and easier process. Long ago, Schrödinger[182] and Löwdin[181] suggested that from the physical point of view, the genetic material could be considered as an "aperiodic solid". Certainly, there were still a lot of molecular engineering and a long road between the "aperiodic solid" and probably "senseless code" of the first RNA molecules and the DNA-enzyme world from the first living cells.
Along the prolonged evolutive scenario, the ancestral ribopolynucleotides would be closed circular structures such as viroids and virusoids. Two types of replication processes could be envisaged for ancient RNA duplexes, both based on the tetris model, namely a self-assembly of monomeric precursor units. In the stem-loop junction, a replicated duplex would be formed by assembly units on an ss-chain template. In the tetris replication model, precursor units recognize complementary H-bonding surface on the major groove of ds-RNA stem. No parental strand separation nor base rotation are required for this model. Interestingly, a convenient base rotation in RNA tetraplex stem would convert the conservative to a semi-conservative replication mode, and if there is no terminal loops, then segregation would result in two free duplexes.
Regarding non-enzymatic RNA replication, although some torsional stress can be released by rotation and rewinding of terminal loops, it is logical to assume that relieving of the accumulated unwinding turns during replication can be also resolved by formation of the RNA tetraplex, This transient structure suggested for releasing the replication-induced torsional stress would be equivalent to the analogous DNA tetraplex and could serve to similar relaxing functions[147,150].
The stereochemistry of the zipper model and ORNAM dumbbell replication is reasonable and uses well-known aspects of RNA conformation and dynamics, but one must be cautious on its precise molecular geometry, since only bibliographic data and computational parameters have been utilized in this work. It must be taken into account the dictum that it is relatively easy to build stereochemically feasible models, difficult to prove that such models are wrong, and even more difficult to prove that they are right. Furthermore, as it is extremely difficult to study and describe molecular processes that occurred about 109 years ago, the proposed RNA formation and replication mode should be treated rather as working and seminal models in which mechanistic aspects have been emphasized.
At the beginning, there was water, CO2, methane, formaldehyde, cyanide, ammonia, and other very simple molecules. Then, at another beginning, suitable heterocycles (nitrogenous bases) and sugars (ribose) were self-assembled along a clay matrix and polymerized into NAs by phosphate links. Then, non-enzymatic self-replication of the simplest RNA specimens began to reproduce them. Until here, these beginnings are only the possible history of small and simple RNAs. However, life is much more than those beginnings: it implicates assembly of amino acids into proteins and enzymes, genetic code, enzymatic NAs replication, energy handling, metabolism, etc. Obviously, that is an important and fascinating scenario of biogenesis that remains to be explored, but it would be another history.
Acknowledgements
The authors are deeply grateful to Abasolo MI, Bella JL, Blanco MM, Blázquez-Castro A, Casas AG, Colombo LL, Durantini EN, Espada J, Felix-Pozzi MN, Gabay O, Herkovits J, Horobin RW, Lu A, Lu G, Romero SA, Solari Abasolo S, Solarz GR, Soltermann A, Stockert A, and Stockert F for valuable collaboration, advices, stimulating discussions, and critical reading of the manuscript.
Authors contribution
The author contributed solely to the article.
Conflicts of interest
The author declares no conflicts of interest.
Ethical approval
Not applicable.
Consent for publication
Not applicable.
Availability of data and materials
All data generated or analyzed in this study are included in the published article.
Funding
None.
Copyright
© The Author(s) 2023.
References
-
1. Bernal JD. The physical basis of life. Proc R Soc A. 1949;62(9):537.[DOI]
-
2. Goldschmidt VM. Geochemical aspects of the origin of complex organic molecules on the Earth, as precursors to organic life. New Biol. 1952;12:97-105.
-
3. Calvin M. The chemistry of life: 3. How life originated on Earth and on the world beyond. Chem Eng News. 1961;39(21):96-104.
-
4. Carvajal G. Bases experimentales sobre el origen de la materia viva. In: Ondarza RN, editor. Introduccion a la biologia moderna. 1964. Mexico: Fondo de Cultura Económico; 1964. p. 13-53.
-
6. Cronin JR. Clues from the origin of the solar system: meteorites. In: Brack A, editor. The molecular origins of life: Assembling pieces of the puzzle. New York: Cambridge University Press; 1998. p. 119-146.[DOI]
-
20. Ball P. A new idea for how to assemble life[Internet]. 2023. Available from: https://www.quantamagazine.org/
-
21. Dauvillier A. The photochemical origin of life. New York: ACS Publications; 1965.
-
22. Aguilera Mochon JA. El origen de la vida: La aparicion de los primeros microorganismos. Barcelona: RBA Libros; 2018. Spanish.
-
26. Guschlbauer W. Nucleic acid structure. New York: Springer; 1976.[DOI]
-
27. Steward PR, Letham DS. The ribonucleic acids. 1st ed. New York: Springer; 1973.[DOI]
-
29. Saenger W. Principles of nucleic acid structure. New York: Springer; 1984.[DOI]
-
30. Blackburn GM, Gait MJ. Nucleic acids in chemistry and biology. Oxford: Oxford University Press; 1990.
-
34. Wei X, Meng D, You C. In vitro metabolic engineering: current status and recent progress. In: Liu L, Du G, Liu Y, editors. Systems and synthetic metabolic engineering. New York: Academic Press; 2020. p. 183-206.[DOI]
-
37. Stockert JC, del Castillo P. Estudios espectroscopicos sobre agregación de cromóforos planos e implicaciones en la síntesis prebiótica de ácidos nucleicos. In: Proceedings of the 3rd Meeting of the Spanish Society for Cell Biology; 1989; Leioa, Spain. 1989. p. 334-335. Spanish.
-
39. Stockert JC, Blazquez-Castro A. Biomedical overview of melanin. 2. Updating molecular modeling, synthesis mechanism, and supramolecular properties regarding melanoma therapy. Biocell. 2022;46(6):1391-1415.[DOI]
-
40. Stockert JC, Espada J, Blazquez-Castro A. Melanin-binding colorants: Updating molecular modeling, staining and labeling mechanisms, and biomedical perspectives. Colorants. 2022;1(1):91-120.[DOI]
-
41. Stockert JC, Felix-Pozzi MN. Updating curvature of graphitic materials: Molecular modeling studies on the spiral structure of graphene and eumelanin models. In: Grant CD, editor. Advances in Biology. New York: Nova Science Publisher; 2022.
-
43. Stockert JC. Stereoscopy of computer-drawn molecular structures. Biochem Educ. 1994;22(1):23-25.[DOI]
-
46. Stockert JC, Blazquez-Castro A. Fluorescence microscopy in life sciences. Sharjah: Bentham Science Publishers; 2017.[DOI]
-
52. Lipmann F. Metabolic generation and utilization of phosphate bond energy. In: Nord FF, Werkman CH, editors. Advances in Enzymology and Related Areas of Molecular Biology.New York: Interscience Publishers; 1941. p. 99-162.[DOI]
-
53. Stryer L. Biochemistry. 3rd ed. San Francisco: W. Freeman; 1988.
-
54. Kulaev IS, Vagabov V, Kulakovskaya T. The biochemistry of inorganic polyphosphates. 2nd ed. Chichester: John Wiley & Son; 2004.
-
56. Mortland MM. Clay-organic complexes and interactions. Adv Agron. 1970;22:75-117.[DOI]
-
57. Sawai H, Orgel LE. Oligonucleotide synthesis catalyzed by the zinc(2+) ion. J Am Chem Soc. 1975;97(12):3532-3533.[DOI]
-
59. Ferris JP. Mineral catalysis and prebiotic synthesis: Montmorillonite-catalyzed formation of RNA. Elements. 2005;1(3):145-3149.[DOI]
-
61. Ertem G, Ferris JP. Sequence- and regio-selectivity in the montmorillonite catalyzed synthesis of RNA. Orig Life Evol Biosph. 2000;30(5):411-422.[DOI]
-
71. Kirby AJ, Younas M. The reactivity of phosphate esters. Diester hydrolysis. J Chem Soc B. 1970;510-513.[DOI]
-
72. Hendricks SB. Lattice structures of clay minerals and some properties. J Geol. 1942;50(3):276-290. Available from: https://www.jstor.org/stable/30070579
-
73. Thomas JM. Sheet silicate intercalates: new agents for unusual chemical conversions. In: Whittingham MS, Jacobson AJ, editors. Intercalation chemistry. New York: Academic Press; 1982. p. 55-99.
-
74. Hofmann U, Endell K, Wilm D. Kristallstruktur und Quellung von Montmorillonit. Z Kristallogr Cryst Mater. 1933;86:340-348. German.[DOI]
-
75. van Olphen H. An introduction to clay colloidal chemistry. New York: Interscience Publishers; 1977.
-
76. Marco-Brown JL, Arcco MM, Sanchez RMT, dos Santos Afonso M. Adsorption of picloram herbicide on montmorillonite: kinetic and equilibrium studies. Colloids Surf A. 2014;449:121-128.[DOI]
-
77. Yariv S, Russell JD, Farmer VC. Infrared study of the adsorption of benzoic acid and nitrobenzene in montmorillonite. Israel J Chemistry. 1966;4(5-6):201-213.[DOI]
-
78. Whittingham MS. Intercalation chemistry: an introduction. In: Whittingham MS, Jacobson AJ, editors. Intercalation chemistry. New York: Academic Press; 1982. p. 1-18.
-
79. Rausell-Colom JA, Serratosa JM. Reactions of clays with organic substances. In: Newman ACD, editor. Chemistry of clays and clay minerals. New York: Interscience Publishers; 1987. p. 371-422.
-
80. Greenland DJ, Laby RH, Quirk JP. Adsorption of amino-acids and peptides by montmorillonite and illite. Part 2 - Physical adsorption. Trans Faraday Soc. 1965;61:2024-2035.[DOI]
-
81. Greene-Kelly R. Sorption of aromatic organic compounds by montmorillonite. Part 1 - orientation studies. Trans Faraday Soc. 1955;51:412-424.[DOI]
-
82. Lailach GE, Thompson TD, Brindley GW. Absorption of pyrimidines, purines, and nucleosides by Li-, Na-, Mg-, and Ca-montmorillonite (Clay-organic studies XII). Clays Clay Miner. 1968;16(4):285-293.[DOI]
-
83. Lailach GE, Thompson TD, Brindley GW. Absorption of pyrimidines, purines, and nucleosides by Co-, Ni-, Cu-, and Fe(III)-montmorillonite (Clay-organic studies XIII). Clays Clay Miner. 1968;16(4):295-301.[DOI]
-
84. Lailach GE, Brindley GW. Specific co-absorption of purines and pyrimidines by montmorillonite (Clay-organic studies XV). Clays Clay Miner. 1969;17(2):95-100.[DOI]
-
86. Plesa Chicinas R, Stefan R, Maicaneanu A. Ability of a montmorillonitic clay to interact with cationic and anionic dyes in aqueous solutions. J Mol Struct. 2018;1154:187e195.[DOI]
-
87. del Rio MS, Martinetto P, Reyes-Valerio C, Dooryhee E, Suarez M. Synthesis and acid resistance of Maya blue pigment. Archaeometry. 2006;48(1):115-130.[DOI]
-
88. Bujdak J. Effect of the layer charge of clay minerals on optical properties of organic dyes. A review. Appl Clay Sci. 2006;34(1-4):58-73.[DOI]
-
91. Jheeta S, Joshi PC. Prebiotic RNA synthesis by montmorillonite catalysis. Life. 2014;4(3):318-330.[DOI]
-
93. Frank-Kamenetskii MD. Protonated DNA structures. In: Eckstein F, Lilley DMJ, editors. Nucleic acids and molecular biology. Berlin: Springer; 1990. p. 1-15.[DOI]
-
94. Arnott S, Bond PJ, Chandrasekaran R. Visualization of an unwound DNA duplex. Nature. 1980;287(5782):561-563.[DOI]
-
96. Stockert JC. Cytochemistry of nucleic acids: binding mechanisms of dyes and fluorochromes. Microsc Electron Biol Celular. 1985;9(2):89-131.[PubMed]
-
97. Udenfriend S. Fluorescence assay in biology and medicine. New York: Academic Press; 1969.[DOI]
-
99. Low PF. The swelling of clay: III. Dissociation of exchangeable cations. Soil Sci Soc Amer J. 1981;45(6):1074-1078.[DOI]
-
102. van de Sande JH, Ramsing NB, Germann MW, Elhorst W, Kalisch BW, Kitzing EV, et al. Parallel stranded DNA. Science. 1988;241(4865):551-557.[DOI]
-
104. Kawamura K, Okamoto F. Cyclization and dimerization of hexanucleotides containing guanine and cytosine with water-soluble carbodiimide. Viva Origino. 2001;29(4):162-167.[DOI]
-
109. Sheldon CC, Jeffries AC, Davies C, Symons RH. RNA self-cleavage by the hammerhead structure. In: Eckstein F, Lilley DMJ, editors. Nucleic Acids and Molecular Biology 4. Berlin: Springer; 1990. p. 227-242.[DOI]
-
110. Lamond AI, Barabino S, Blencowe BJ. The mammalian pre-mRNA splicing apparatus. In: Eckstein F, Lilley DMJ, editors. Nucleic Acids and Molecular Biology 4. Berlin: Springer; 1990. p. 243-257.[DOI]
-
111. Tinoco I, Puglisi JD, Wyatt JR. RNA folding. In: Eckstein F, Lilley DMJ, editors. Nucleic Acids and Molecular Biology 4. Berlin: Springer; 1990. p. 205-226.[DOI]
-
113. Schimmel P. Interpretation of experiments that delineate transfer RNA recognition in vivo and in vitro. In: Eckstein F, Lilley DMJ, editors. Nucleic Acids and Molecular Biology 4. Berlin: Springer; 1990. p. 274-287.[DOI]
-
121. Keese P, Symons RH. The structure of viroids and virusoids. In: Semancik JS, editor. Viroids and viroid-like pathogens. Florida: CRC; 1987. p. 1-47.
-
122. Flores R, Di Serio F, Hernandez C. Viroids: the noncoding genomes. Sem Virology. 1997;8(1):65-73.[DOI]
-
132. Thiele D, Guschlbauer W, Favre A. Protonated polynucleotide structures. X. Optical properties of poly(I)-poly(C) and its disproportionation complexes. Biochim Biophys Acta. 1972;272(1):22-26.[PubMed]
-
142. Simundza G, Sakore TD, Sobell HM. Base-pairing configurations between purines and pyrimidines in the solid state. V. Crystal and molecular structure of two 1:1 hydrogen-bonded complexes, 1-methyluracil: 9-ethyl-8-bromo-2,6-diaminopurine and 1-ethylthymine: 9-ethyl-8-bromo-2,6-diaminopurine. J Mol Biol. 1970;48(2):263-278.
-
147. Morgan AR. Do multistranded polynucleotides have a biological role? Trends Biochem Sci. 1979;4(10):244-247.[DOI]
-
148. Stockert JC. Stereochemistry of general genetic recombination: Paranemic duplexes and duplex-tetraplex junctions as possible DNA structures for homologous pairing, strand exchange and hybrid resolution. Biol Zentbl. 1992;111:150-163. Available from: https://www.cabidigitallibrary.org/doi/full/10.5555/19920100701
-
153. Stockert JC. Pairing of underwound DNA duplexes as hypothetical intermediates in genetic recombination. Naturwissenschaften. 1986;73(2):100-101.[PubMed]
-
156. McGavin S. Relationships and transformations between some nucleic acid models. J Theor Biol. 1980;85(4):665-672.[DOI]
-
157. McGavin S, Wilson HR, Barr GC. Intercalated nucleic acid double helices: a stereochemical possibility. J Mol Biol. 1966;22(1):187-191.[DOI]
-
158. Zimmerman SB. X-ray study by fiber diffraction methods of a self-aggregate of guanosine-5'-phosphate with the same helical parameters as poly(rG). J Mol Biol. 1976;106(3):663-672.[PubMed]
-
159. Neidle S. Human telomeric G-quadruplex: the current status of telomeric G-quadruplexes as therapeutic targets in human cancer. FEBS J. 2010;277(5):1118-1125.[PubMed]
-
160. Kim SH. Three-dimensional structure of transfer RNA. Prog Nucleic Acid Res Mol Biol. 1976;17:181-216.[DOI]
-
161. Sequin U. How to view stereoscopic pictures of crystal structures and molecular models. Experientia. 1977;33(8):1115-1118.[DOI]
-
162. Blackburn EH. Structure and function of telomeres. Nature. 1991;350(6319):569-573.[PubMed]
-
165. Phan AT, Gueron M, Leroy JL. The solution structure and internal motions of a fragment of the cytidine-rich strand of the human telomere. J Mol Biol. 2000;299(1):123-144.[PubMed]
-
167. Kitaigorodski AI. Order and disorder in the world of atoms. New York: Springer; 1967.
-
168. Nagl W. Molecular phylogeny. In: Raup DM, Jablonski D, editors. Patterns and processes in the history of life. Berlin: Springer; 1986. p. 223-232.[DOI]
-
170. Scott WG. Molecular palaeontology: understanding catalytic mechanisms in the RNA world by excavating clues from a ribozyme three-dimensional structure. Biochem Soc Trans. 1996;24(3):604-608.[PubMed]
-
171. Moody ERR, Alvarez-Carretero S, Mahendrarajah TA, Clark JW, Betts HC, Dombrowski N, et al. The nature of the last universal common ancestor and its impact on the early Earth system. Nat Ecol Evol. 2024;8(9):1654-1666.
-
172. Cech TR, Bass BL. Biological catalysis by RNA. Annu Rev Biochem. 1986;55:599-629.[PubMed]
-
180. Fong P. The replication of the DNA molecule. Proc Natl Acad Sci USA. 1964;52(3):641-647.[DOI]
-
181. Lowdin PO. Quantum genetics and the aperiodic solid: some aspects on the biological problems of heredity, mutations, aging, and tumors in view of the quantum theory of the DNA molecule. Adv Quant Chem. 1966;2:213-360.[DOI]
-
182. Schrodinger E. What is life? The physical aspect of the living cell. New York: Cambridge Univ. Press; 1944.
Copyright
© The Author(s) 2023. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Publisher’s Note
Share And Cite




